Difference between revisions of "YtsJ"
| Line 143: | Line 143: | ||
=References= | =References= | ||
| − | + | <pubmed>22740702</pubmed> | |
| − | <pubmed>16479537, 16788182 ,12949160 9387221 | + | <pubmed>16479537, 16788182 ,12949160 9387221</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 16:27, 30 October 2012
- Description: malic enzyme, forms a transhydrogenation cycle with MalS for balancing of NADPH
| Gene name | ytsJ |
| Synonyms | |
| Essential | no |
| Product | NADP-dependent malate dehydrogenase |
| Function | malate utilization |
| Gene expression levels in SubtiExpress: ytsJ | |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 43 kDa, 5.046 |
| Gene length, protein length | 1230 bp, 410 aa |
| Immediate neighbours | accD, dnaE |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 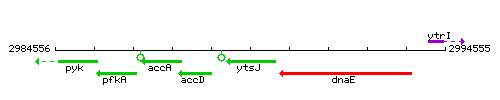
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed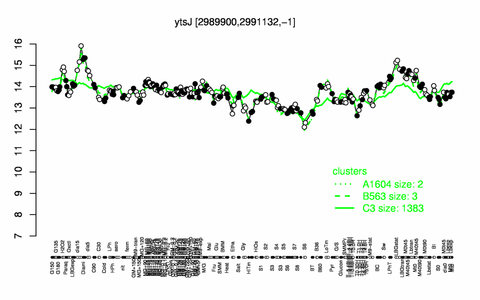
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU29220
Phenotypes of a mutant
Poor growth with malate as single carbon source PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: (S)-malate + NADP+ = pyruvate + CO2 + NADPH (according to Swiss-Prot) malate <--> pyruvate
- Protein family: malic enzymes family (according to Swiss-Prot)
- Paralogous protein(s): MleA
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
Database entries
- Structure: 1WW8 (from Pyrococcus horikoshii, 46% identity, 63% similarity)
- UniProt: O34962
- KEGG entry: [3]
- E.C. number: 1.1.1.38
Additional information
Expression and regulation
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- GP612 (spc), available in Jörg Stülke's lab
- GP1143 (spc), available in Jörg Stülke's lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- FLAG-tag construct: GP1131 (spc, based on pGP1331), available in Jörg Stülke's lab
- Antibody:
Labs working on this gene/protein
Stephane Aymerich, Microbiology and Molecular Genetics, INRA Paris-Grignon, France
Your additional remarks
References