Difference between revisions of "RnhB"
m (Reverted edits by 134.76.70.252 (talk) to last revision by Jstuelk) |
|||
| Line 100: | Line 100: | ||
* '''Operon:''' | * '''Operon:''' | ||
| − | * '''[ | + | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=rnhB_1677451_1678218_1 rnhB] {{PubMed|22383849}} |
| + | |||
| + | * '''Sigma factor:''' | ||
* '''Regulation:''' | * '''Regulation:''' | ||
| Line 106: | Line 108: | ||
* '''Regulatory mechanism:''' | * '''Regulatory mechanism:''' | ||
| − | * '''Additional information:''' | + | * '''Additional information:''' |
=Biological materials = | =Biological materials = | ||
Revision as of 08:55, 13 April 2012
- Description: RNase HII, endoribonuclease
| Gene name | rnhB |
| Synonyms | rnh |
| Essential | no |
| Product | Mn2+-dependent RNase HII |
| Function | endonucleolytic cleavage of RNA in RNA-DNA hybrid molecules |
| MW, pI | 28 kDa, 5.518 |
| Gene length, protein length | 765 bp, 255 aa |
| Immediate neighbours | rbgA, ylqG |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 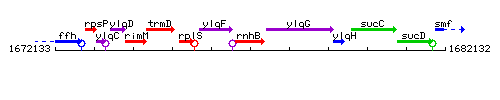
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU16060
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Endonucleolytic cleavage to 5'-phosphomonoester (according to Swiss-Prot)
- Protein family: RNase HII family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- cytoplasm (according to Swiss-Prot)
Database entries
- Structure:
- UniProt: O31744
- KEGG entry: [2]
- E.C. number: 3.1.26.4.
Additional information
Expression and regulation
- Operon:
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications
Sanae Fukushima, Mitsuhiro Itaya, Hiroaki Kato, Naotake Ogasawara, Hirofumi Yoshikawa
Reassessment of the in vivo functions of DNA polymerase I and RNase H in bacterial cell growth.
J Bacteriol: 2007, 189(23);8575-83
[PubMed:17905985]
[WorldCat.org]
[DOI]
(I p)
Mitsuru Haruki, Yasuo Tsunaka, Masaaki Morikawa, Shigenori Kanaya
Cleavage of a DNA-RNA-DNA/DNA chimeric substrate containing a single ribonucleotide at the DNA-RNA junction with prokaryotic RNases HII.
FEBS Lett: 2002, 531(2);204-8
[PubMed:12417313]
[WorldCat.org]
[DOI]
(P p)
M Itaya, A Omori, S Kanaya, R J Crouch, T Tanaka, K Kondo
Isolation of RNase H genes that are essential for growth of Bacillus subtilis 168.
J Bacteriol: 1999, 181(7);2118-23
[PubMed:10094689]
[WorldCat.org]
[DOI]
(P p)
N Ohtani, M Haruki, M Morikawa, R J Crouch, M Itaya, S Kanaya
Identification of the genes encoding Mn2+-dependent RNase HII and Mg2+-dependent RNase HIII from Bacillus subtilis: classification of RNases H into three families.
Biochemistry: 1999, 38(2);605-18
[PubMed:9888800]
[WorldCat.org]
[DOI]
(P p)