Difference between revisions of "AmyE"
m (Reverted edits by 134.76.70.252 (talk) to last revision by Jstuelk) |
|||
| Line 101: | Line 101: | ||
* '''Operon:''' ''[[amyE]]'' {{PubMed|11320136}} | * '''Operon:''' ''[[amyE]]'' {{PubMed|11320136}} | ||
| − | * '''[ | + | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=amyE_327618_329597_1 amyE] {{PubMed|22383849}} |
| + | |||
| + | * '''Sigma factor:''' [[SigA]] {{PubMed|3123701}} | ||
* '''Regulation:''' | * '''Regulation:''' | ||
Revision as of 13:55, 12 April 2012
- Description: alpha-amylase
| Gene name | amyE |
| Synonyms | amyA |
| Essential | no |
| Product | alpha-amylase) |
| Function | starch degradation |
| Metabolic function and regulation of this protein in SubtiPathways: Sugar catabolism | |
| MW, pI | 72 kDa, 5.85 |
| Gene length, protein length | 1980 bp, 660 aa |
| Immediate neighbours | ycgB, ldh |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 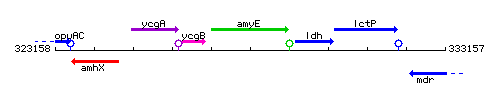
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU03040
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Endohydrolysis of (1->4)-alpha-D-glucosidic linkages in oligosaccharides and polysaccharides (according to Swiss-Prot)
- Protein family: glycosyl hydrolase 13 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: secreted (according to Swiss-Prot), extracellular (signal peptide) PubMed
Database entries
- Structure: 1BAG (complex with maltopentaose)
- UniProt: P00691
- KEGG entry: [3]
- E.C. number: 3.2.1.1
Additional information
Expression and regulation
- Additional information:
Biological materials
- Mutant: GP550 (cat), available in Stülke lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Additional publications: PubMed