Difference between revisions of "GltA"
m (Reverted edits by 134.76.70.252 (talk) to last revision by Jstuelk) |
|||
| Line 44: | Line 44: | ||
= This gene is a member of the following [[regulons]] = | = This gene is a member of the following [[regulons]] = | ||
{{SubtiWiki regulon|[[GltC regulon]]}}, | {{SubtiWiki regulon|[[GltC regulon]]}}, | ||
| + | {{SubtiWiki regulon|[[FsrA regulon]]}}, | ||
{{SubtiWiki regulon|[[TnrA regulon]]}} | {{SubtiWiki regulon|[[TnrA regulon]]}} | ||
| Line 122: | Line 123: | ||
** expressed in the presence of ammonium {{PubMed|11029411}} | ** expressed in the presence of ammonium {{PubMed|11029411}} | ||
** repressed in the absence of good nitrogen sources (glutamine or ammonium) ([[TnrA]]) {{PubMed|11029411}} | ** repressed in the absence of good nitrogen sources (glutamine or ammonium) ([[TnrA]]) {{PubMed|11029411}} | ||
| + | ** part of the iron sparing response, strong down-regulation in a'' [[fur]]'' mutant ([[Fur]], [[FsrA]]) {{PubMed|22389480}} | ||
* '''Regulatory mechanism:''' | * '''Regulatory mechanism:''' | ||
** [[GltC]]: transcription activation {{PubMed|2548995}} | ** [[GltC]]: transcription activation {{PubMed|2548995}} | ||
** [[TnrA]]: transcription repression {{PubMed|11029411}} | ** [[TnrA]]: transcription repression {{PubMed|11029411}} | ||
| + | ** [[Fur]]: indirect effect via control of ''[[fsrA]]'' expression {{PubMed|22389480}} | ||
| + | ** [[FsrA]]: translation inhibition {{PubMed|22389480}} | ||
* '''Additional information:''' subject to Clp-dependent proteolysis upon glucose starvation {{PubMed|17981983}} | * '''Additional information:''' subject to Clp-dependent proteolysis upon glucose starvation {{PubMed|17981983}} | ||
| Line 159: | Line 163: | ||
<pubmed>16143852 12859215</pubmed> | <pubmed>16143852 12859215</pubmed> | ||
==Original publications== | ==Original publications== | ||
| − | <pubmed>12823818,18199747 ,11029411,12850135 7559360 15150225 2548995 17183217, 17608797,17134717,14523131,12823818, 18326565,17981983,18763711 20933603 17012385 </pubmed> | + | <pubmed>12823818,18199747 ,11029411,12850135 7559360 15150225 2548995 17183217, 17608797,17134717,14523131,12823818, 18326565,17981983,18763711 20933603 17012385 22389480</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 18:03, 14 March 2012
- Description: large subunit of glutamate synthase
| Gene name | gltA |
| Synonyms | |
| Essential | no |
| Product | glutamate synthase (large subunit) |
| Function | glutamate biosynthesis |
| Interactions involving this protein in SubtInteract: GltA | |
| Metabolic function and regulation of this protein in SubtiPathways: Ammonium/ glutamate | |
| MW, pI | 168 kDa, 5.47 |
| Gene length, protein length | 4560 bp, 1520 amino acids |
| Immediate neighbours | gltB, gltC |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 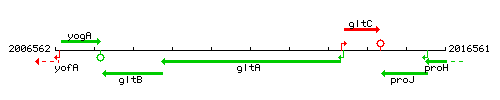
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of amino acids, membrane proteins
This gene is a member of the following regulons
GltC regulon, FsrA regulon, TnrA regulon
The gene
Basic information
- Locus tag: BSU18450
Phenotypes of a mutant
auxotrophic for glutamate
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: 2 L-glutamate + NADP+ = L-glutamine + 2-oxoglutarate + NADPH (according to Swiss-Prot) 2 L-glutamate + NADP(+) <=> L-glutamine + 2-oxoglutarate + NADPH
- Protein family: glutamate synthase family (according to Swiss-Prot) glutamate synthase family
- Paralogous protein(s): YerD
Extended information on the protein
- Kinetic information:
- Domains:
- Glutamine amidotransferase type-2 domain (22-415)
- Nucleotide binding domain (1060-1112)
- Modification:
- Cofactor(s): 3Fe-4S, FAD, FMN
- Effectors of protein activity:
- Localization: membrane associated PubMed, cytoplasm
Database entries
- UniProt: P39812
- KEGG entry: [3]
- E.C. number: 1.4.1.13 3 1.4.1.13]
Additional information
subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Regulation:
- expression activated by glucose (11 fold) (CcpA, GltC) PubMed
- repressed by arginine (GltC, RocG) PubMed
- expressed in the presence of ammonium PubMed
- repressed in the absence of good nitrogen sources (glutamine or ammonium) (TnrA) PubMed
- part of the iron sparing response, strong down-regulation in a fur mutant (Fur, FsrA) PubMed
- Regulatory mechanism:
- Additional information: subject to Clp-dependent proteolysis upon glucose starvation PubMed
Biological materials
- Mutant: GP807 (del gltAB::tet), GP222 (gltA under the control of p-xyl), available in Stülke lab
- Expression vector:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody:
Labs working on this gene/protein
Linc Sonenshein, Tufts University, Boston, MA, USA Homepage
Jörg Stülke, University of Göttingen, Germany Homepage
Your additional remarks
References
Reviews
Original publications