Difference between revisions of "Spo0E"
| Line 1: | Line 1: | ||
| − | * '''Description:''' [[Spo0A]]-P phosphatase <br/><br/> | + | * '''Description:''' [[Spo0A]]-P phosphatase, control of the [[phosphorelay]] <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
Revision as of 09:31, 28 November 2009
- Description: Spo0A-P phosphatase, control of the phosphorelay
| Gene name | spo0E |
| Synonyms | |
| Essential | no |
| Product | Spo0A-P phosphatase |
| Function | initiation of sporulation |
| MW, pI | 9 kDa, 6.483 |
| Gene length, protein length | 255 bp, 85 aa |
| Immediate neighbours | ykvA, eag |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 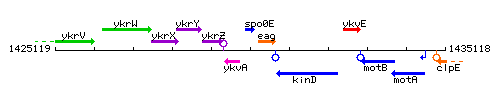
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU13640
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: spo0E family (according to Swiss-Prot)
- Paralogous protein(s): YnzD
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
Database entries
- Structure:
- UniProt: P05043
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: spo0E PubMed
- Additional information: Spo0E is degraded by FtsH
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Tony Wilkinson, York University, U.K. homepage
Your additional remarks
References
Ai Thi Thuy Le, Wolfgang Schumann
The Spo0E phosphatase of Bacillus subtilis is a substrate of the FtsH metalloprotease.
Microbiology (Reading): 2009, 155(Pt 4);1122-1132
[PubMed:19332814]
[WorldCat.org]
[DOI]
(P p)
Andrea Feucht, Laura Abbotts, Jeffery Errington
The cell differentiation protein SpoIIE contains a regulatory site that controls its phosphatase activity in response to asymmetric septation.
Mol Microbiol: 2002, 45(4);1119-30
[PubMed:12180929]
[WorldCat.org]
[DOI]
(P p)
Sophie J Stephenson, Marta Perego
Interaction surface of the Spo0A response regulator with the Spo0E phosphatase.
Mol Microbiol: 2002, 44(6);1455-67
[PubMed:12067336]
[WorldCat.org]
[DOI]
(P p)
I Lucet, A Feucht, M D Yudkin, J Errington
Direct interaction between the cell division protein FtsZ and the cell differentiation protein SpoIIE.
EMBO J: 2000, 19(7);1467-75
[PubMed:10747015]
[WorldCat.org]
[DOI]
(P p)
K L Ohlsen, J K Grimsley, J A Hoch
Deactivation of the sporulation transcription factor Spo0A by the Spo0E protein phosphatase.
Proc Natl Acad Sci U S A: 1994, 91(5);1756-60
[PubMed:8127878]
[WorldCat.org]
[DOI]
(P p)
M Perego, J A Hoch
Negative regulation of Bacillus subtilis sporulation by the spo0E gene product.
J Bacteriol: 1991, 173(8);2514-20
[PubMed:1901567]
[WorldCat.org]
[DOI]
(P p)
U Bai, M Lewandoski, E Dubnau, I Smith
Temporal regulation of the Bacillus subtilis early sporulation gene spo0F.
J Bacteriol: 1990, 172(9);5432-9
[PubMed:2118512]
[WorldCat.org]
[DOI]
(P p)
M A Strauch, G B Spiegelman, M Perego, W C Johnson, D Burbulys, J A Hoch
The transition state transcription regulator abrB of Bacillus subtilis is a DNA binding protein.
EMBO J: 1989, 8(5);1615-21
[PubMed:2504584]
[WorldCat.org]
[DOI]
(P p)
M Perego, J A Hoch
Isolation and sequence of the spo0E gene: its role in initiation of sporulation in Bacillus subtilis.
Mol Microbiol: 1987, 1(1);125-32
[PubMed:2838724]
[WorldCat.org]
[DOI]
(P p)