Difference between revisions of "OxdC"
| Line 80: | Line 80: | ||
* '''Swiss prot entry:''' [http://www.uniprot.org/uniprot/O34714 O34714] | * '''Swiss prot entry:''' [http://www.uniprot.org/uniprot/O34714 O34714] | ||
| − | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu | + | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu:BSU33240] |
* '''E.C. number:''' [http://www.expasy.org/enzyme/4.1.1.2 4.1.1.2] | * '''E.C. number:''' [http://www.expasy.org/enzyme/4.1.1.2 4.1.1.2] | ||
Revision as of 03:33, 25 June 2009
- Description: oxalate decarboxylase
| Gene name | oxdC |
| Synonyms | yvrK |
| Essential | no |
| Product | oxalate decarboxylase |
| Function | unknown |
| MW, pI | 43 kDa, 5.101 |
| Gene length, protein length | 1155 bp, 385 aa |
| Immediate neighbours | yvrJ, yvrL |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 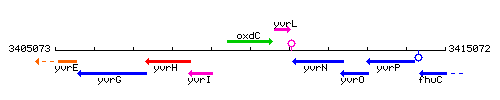
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU33240
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Oxalate = formate + CO2 (according to Swiss-Prot)
- Protein family:
- Paralogous protein(s): OxdD
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s): requires Mn(II) for activity PubMed
- Effectors of protein activity:
- Interactions:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- Structure: 1UW8
- Swiss prot entry: O34714
- KEGG entry: [2]
- E.C. number: 4.1.1.2
Additional information
Expression and regulation
- Regulation: induced by acidic growth conditions PubMed
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
John Helmann, Cornell University, USA Homepage
Your additional remarks
References