Difference between revisions of "Sandbox"
| Line 1: | Line 1: | ||
| − | * '''Description:''' | + | * '''Description:''' cystathionine b-lyase <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Gene name''' | |style="background:#ABCDEF;" align="center"|'''Gene name''' | ||
| − | |'' | + | |''metC'' |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Synonyms''' || '' | + | |style="background:#ABCDEF;" align="center"| '''Synonyms''' || ''yjcJ'' |
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''Essential''' || no | |style="background:#ABCDEF;" align="center"| '''Essential''' || no | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Product''' || | + | |style="background:#ABCDEF;" align="center"| '''Product''' || cystathionine b-lyase |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Function''' || | + | |style="background:#ABCDEF;" align="center"|'''Function''' || biosynthesis of methionine |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || | + | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 42 kDa, 5.757 |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || | + | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || 1170 bp, 390 aa |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[ | + | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[metI]]'', ''[[yjcK]]'' |
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+[EMBLCDS: | + | |colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+[EMBLCDS:CAB13045]+-newId sequences] <br/> (Barbe ''et al.'', 2009)''' |
|- | |- | ||
| − | |colspan="2" | '''Genetic context''' <br/> [[Image: | + | |- |
| + | |- | ||
| + | |colspan="2" | '''Genetic context''' <br/> [[Image:yjcJ_context.gif]] | ||
| + | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
|- | |- | ||
|} | |} | ||
| Line 28: | Line 31: | ||
__TOC__ | __TOC__ | ||
| − | <br/><br/> | + | <br/><br/><br/><br/> |
=The gene= | =The gene= | ||
| Line 34: | Line 37: | ||
=== Basic information === | === Basic information === | ||
| − | * '''Locus tag:''' | + | * '''Locus tag:''' BSU11880 |
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
| Line 40: | Line 43: | ||
=== Database entries === | === Database entries === | ||
| − | * '''DBTBS entry:''' | + | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/yjcIJ.html] |
| − | * '''SubtiList entry:''' [http://genolist.pasteur.fr/SubtiList/genome.cgi?gene_detail+ | + | * '''SubtiList entry:''' [http://genolist.pasteur.fr/SubtiList/genome.cgi?gene_detail+BG13163] |
=== Additional information=== | === Additional information=== | ||
| Line 51: | Line 54: | ||
=== Basic information/ Evolution === | === Basic information/ Evolution === | ||
| − | * '''Catalyzed reaction/ biological activity:''' | + | * '''Catalyzed reaction/ biological activity:''' L-cystathionine + H<sub>2</sub>O = L-homocysteine + NH<sub>3</sub> + pyruvate (according to Swiss-Prot) |
| − | * '''Protein family:''' | + | * '''Protein family:''' trans-sulfuration enzymes family (according to Swiss-Prot) |
* '''Paralogous protein(s):''' | * '''Paralogous protein(s):''' | ||
| Line 71: | Line 74: | ||
* '''Interactions:''' | * '''Interactions:''' | ||
| − | * '''Localization:''' | + | * '''Localization:''' cytoplasm (according to Swiss-Prot) |
=== Database entries === | === Database entries === | ||
| Line 77: | Line 80: | ||
* '''Structure:''' | * '''Structure:''' | ||
| − | * '''Swiss prot entry:''' [http://www.uniprot.org/uniprot/ | + | * '''Swiss prot entry:''' [http://www.uniprot.org/uniprot/O31632 O31632] |
| − | * '''KEGG entry:''' | + | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu+BSU11880] |
| − | * '''E.C. number:''' | + | * '''E.C. number:''' [http://www.expasy.org/enzyme/4.4.1.8 4.4.1.8] |
=== Additional information=== | === Additional information=== | ||
| Line 87: | Line 90: | ||
=Expression and regulation= | =Expression and regulation= | ||
| − | * '''Operon:''' ''[[ | + | * '''Operon:''' ''[[metI]]-[[metC]]'' [http://www.ncbi.nlm.nih.gov/sites/entrez/11832514 PubMed] |
| − | * ''' | + | * '''Sigma factor:''' [[SigA]] [http://www.ncbi.nlm.nih.gov/sites/entrez/11832514 PubMed] |
| − | * '''Regulation:''' | + | * '''Regulation:''' |
| + | **repressed by casamino acids [http://www.ncbi.nlm.nih.gov/pubmed/12107147 PubMed] | ||
| + | ** induced by methionine starvation ([[S-box]]) [http://www.ncbi.nlm.nih.gov/sites/entrez/10094622 PubMed] | ||
| − | * '''Regulatory mechanism:''' | + | * '''Regulatory mechanism:''' [[S-box]]: transcription termination/ antitermination, the [[S-box]] [[riboswitch]] binds S-adenosylmethionine resulting in termination [http://www.ncbi.nlm.nih.gov/sites/entrez/10094622 PubMed] |
* '''Additional information:''' | * '''Additional information:''' | ||
| Line 99: | Line 104: | ||
=Biological materials = | =Biological materials = | ||
| − | * '''Mutant:''' | + | * '''Mutant:''' |
* '''Expression vector:''' | * '''Expression vector:''' | ||
| Line 117: | Line 122: | ||
=References= | =References= | ||
| − | <pubmed> | + | <pubmed>12107147, </pubmed> |
| − | # | + | # Mäder et al. (2002) Transcriptome and Proteome Analysis of ''Bacillus subtilis'' Gene Expression Modulated by Amino Acid Availability. ''J. Bacteriol'' '''184:''' 1844288-4295 [http://www.ncbi.nlm.nih.gov/pubmed/12107147 PubMed] |
| − | |||
# Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | # Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | ||
Revision as of 13:24, 8 June 2009
- Description: cystathionine b-lyase
| Gene name | metC |
| Synonyms | yjcJ |
| Essential | no |
| Product | cystathionine b-lyase |
| Function | biosynthesis of methionine |
| MW, pI | 42 kDa, 5.757 |
| Gene length, protein length | 1170 bp, 390 aa |
| Immediate neighbours | metI, yjcK |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 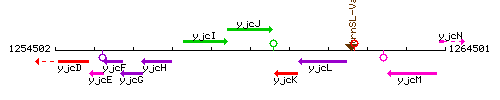
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU11880
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: L-cystathionine + H2O = L-homocysteine + NH3 + pyruvate (according to Swiss-Prot)
- Protein family: trans-sulfuration enzymes family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- Structure:
- Swiss prot entry: O31632
- KEGG entry: [3]
- E.C. number: 4.4.1.8
Additional information
Expression and regulation
- Regulatory mechanism: S-box: transcription termination/ antitermination, the S-box riboswitch binds S-adenosylmethionine resulting in termination PubMed
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Ulrike Mäder, Georg Homuth, Christian Scharf, Knut Büttner, Rüdiger Bode, Michael Hecker
Transcriptome and proteome analysis of Bacillus subtilis gene expression modulated by amino acid availability.
J Bacteriol: 2002, 184(15);4288-95
[PubMed:12107147]
[WorldCat.org]
[DOI]
(P p)