Difference between revisions of "GcvPB"
| Line 118: | Line 118: | ||
=References= | =References= | ||
| + | <pubmed>12107147 15472076, </pubmed> | ||
# Mäder et al. (2002) Transcriptome and Proteome Analysis of ''Bacillus subtilis'' Gene Expression Modulated by Amino Acid Availability. ''J. Bacteriol'' '''184:''' 1844288-4295 [http://www.ncbi.nlm.nih.gov/pubmed/12107147 PubMed] | # Mäder et al. (2002) Transcriptome and Proteome Analysis of ''Bacillus subtilis'' Gene Expression Modulated by Amino Acid Availability. ''J. Bacteriol'' '''184:''' 1844288-4295 [http://www.ncbi.nlm.nih.gov/pubmed/12107147 PubMed] | ||
# Mandal M, Lee M, Barrick JE, Weinberg Z, Emilsson GM, Ruzzo WL, Breaker RR (2004) A glycine-dependent riboswitch that uses cooperative binding to control gene expression. ''Science'' '''306:''' 275-279. [http://www.ncbi.nlm.nih.gov/sites/entrez/15472076 PubMed] | # Mandal M, Lee M, Barrick JE, Weinberg Z, Emilsson GM, Ruzzo WL, Breaker RR (2004) A glycine-dependent riboswitch that uses cooperative binding to control gene expression. ''Science'' '''306:''' 275-279. [http://www.ncbi.nlm.nih.gov/sites/entrez/15472076 PubMed] | ||
# Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | # Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | ||
Revision as of 19:09, 8 June 2009
- Description: glycine decarboxylase (subunit 2)
| Gene name | gcvPB |
| Synonyms | yqhK, gcvP |
| Essential | no |
| Product | glycine decarboxylase (subunit 2) |
| Function | glycine utilization |
| MW, pI | 54 kDa, 5.283 |
| Gene length, protein length | 1464 bp, 488 aa |
| Immediate neighbours | yqhL, gcvPA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 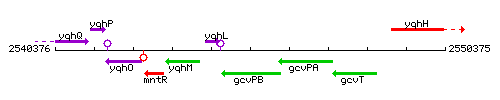
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU24550
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Glycine + H-protein-lipoyllysine = H-protein-S-aminomethyldihydrolipoyllysine + CO2 (according to Swiss-Prot)
- Protein family: C-terminal subunit subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: membrane (according to Swiss-Prot)
Database entries
- Structure:
- Swiss prot entry: P54377
- KEGG entry: [3]
- E.C. number: 1.4.4.2
Additional information
Expression and regulation
- Regulatory mechanism: induction requires a co-operatively acting riboswitch PubMed
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
- Mäder et al. (2002) Transcriptome and Proteome Analysis of Bacillus subtilis Gene Expression Modulated by Amino Acid Availability. J. Bacteriol 184: 1844288-4295 PubMed
- Mandal M, Lee M, Barrick JE, Weinberg Z, Emilsson GM, Ruzzo WL, Breaker RR (2004) A glycine-dependent riboswitch that uses cooperative binding to control gene expression. Science 306: 275-279. PubMed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed