Difference between revisions of "DppA"
| Line 12: | Line 12: | ||
|style="background:#ABCDEF;" align="center"| '''Product''' || D-alanyl-aminopeptidase | |style="background:#ABCDEF;" align="center"| '''Product''' || D-alanyl-aminopeptidase | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Function''' || | + | |style="background:#ABCDEF;" align="center"|'''Function''' || degradation of cell wall peptides |
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 30 kDa, 5.19 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 30 kDa, 5.19 | ||
Revision as of 11:15, 8 April 2009
- Description: D-alanyl-aminopeptidase
| Gene name | dppA |
| Synonyms | dciAA |
| Essential | no |
| Product | D-alanyl-aminopeptidase |
| Function | degradation of cell wall peptides |
| MW, pI | 30 kDa, 5.19 |
| Gene length, protein length | 822 bp, 274 aa |
| Immediate neighbours | proG, dppB |
| Gene sequence (+200bp) | Protein sequence |
| Caution: The sequence for this gene in SubtiList contains errors | |
Genetic context 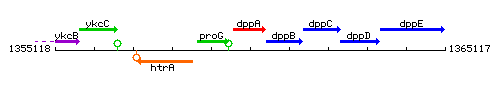
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Coordinates:
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification: phosphorylated on ser/ thr/ tyr PubMed
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure:
- Swiss prot entry:
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon:
- Regulation: repressed by glucose (2.9-fold) PubMed
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
- Blencke et al. (2003) Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways. Metab Eng. 5: 133-149 PubMed
- Lévine et al. (2006) Analysis of the dynamic Bacillus subtilis Ser/Thr/Tyr phosphoproteome implicated in a wide variety of cellular processes. Proteomics 6: 2157-2173 PubMed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed