Difference between revisions of "PdhA"
(→Reviews) |
|||
| Line 159: | Line 159: | ||
=References= | =References= | ||
==Reviews== | ==Reviews== | ||
| − | <pubmed> 19476487 9655937 2227213 6805383 </pubmed> | + | <pubmed> 19476487 9655937 2227213 6805383 24798336 </pubmed> |
| + | |||
==Original publications== | ==Original publications== | ||
<pubmed>9352926, 20525796, 12850135 6414463 11976308 20081037 15378759 24825009</pubmed> | <pubmed>9352926, 20525796, 12850135 6414463 11976308 20081037 15378759 24825009</pubmed> | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 13:51, 28 May 2014
- Description: pyruvate dehydrogenase (E1 alpha subunit), required for Z-ring assembly in a pyruvate-dependent manner
| Gene name | pdhA |
| Synonyms | aceA |
| Essential | yes PubMed, no PubMed |
| Product | pyruvate dehydrogenase (E1 alpha subunit) |
| Function | links glycolysis and TCA cycle |
| Gene expression levels in SubtiExpress: pdhA | |
| Interactions involving this protein in SubtInteract: PdhA | |
| Metabolic function and regulation of this protein in SubtiPathways: pdhA | |
| MW, pI | 41 kDa, 5.837 |
| Gene length, protein length | 1113 bp, 371 aa |
| Immediate neighbours | ykyA, pdhB |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 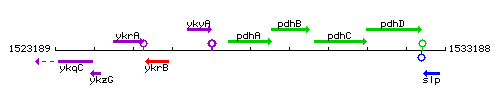
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed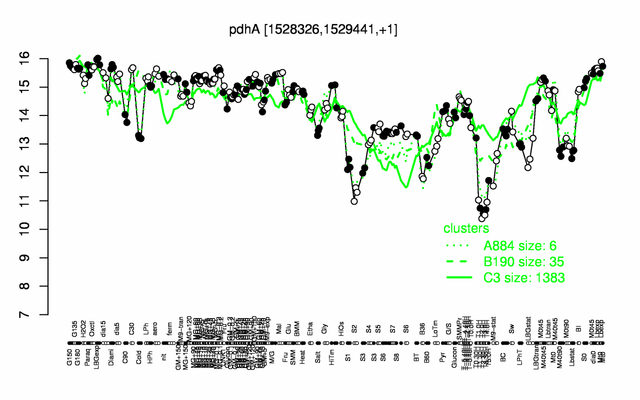
| |
Contents
Categories containing this gene/protein
carbon core metabolism, essential genes, most abundant proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU14580
Phenotypes of a mutant
- pdhA is essential according to Kobayashi et al. PubMed
- the mutant grows slowly but is viable PubMed
- depletion of pdhA and deletion of ezrA have a strong synthetic defect in cell division PubMed
Database entries
- BsubCyc: BSU14580
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Pyruvate + [dihydrolipoyllysine-residue acetyltransferase] lipoyllysine = [dihydrolipoyllysine-residue acetyltransferase] S-acetyldihydrolipoyllysine + CO2 (according to Swiss-Prot)
- Protein family:
Extended information on the protein
- Kinetic information: Michaelis-Menten PubMed
- Modification:
- Cofactors:
- thiamine pyrophosphate
- Effectors of protein activity:
- Inhibited thiamine 2-thiothiazolone diphosphate and NADH PubMed
- Low sensibility to NADPH
- Localization:
- colocalizes with the nucleoid (depending on the availability of pyruvate) PubMed
Database entries
- BsubCyc: BSU14580
- Structure: 1W88 (E1 in complex with subunit binding domain of E2, Geobacillus stearothermophilus)
- UniProt: P21881
- KEGG entry: [3]
- E.C. number: 1.2.4.1
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- stringent response: due to presence of guanine at +1 position of the transcript PubMed
- Additional information:
- The mRNA has a long 5' leader region. This may indicate RNA-based regulation PubMed
- belongs to the 100 most abundant proteins PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium): 5117 PubMed
- number of protein molecules per cell (complex medium with amino acids, without glucose): 18311 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, exponential phase): 4425 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, early stationary phase after glucose exhaustion): 5452 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, late stationary phase after glucose exhaustion): 7055 PubMed
Biological materials
- Mutant:
- Expression vector:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Arthur Aronson, Purdue University, West Lafayette, USA homepage
Your additional remarks
References
Reviews
Mulchand S Patel, Natalia S Nemeria, William Furey, Frank Jordan
The pyruvate dehydrogenase complexes: structure-based function and regulation.
J Biol Chem: 2014, 289(24);16615-23
[PubMed:24798336]
[WorldCat.org]
[DOI]
(I p)
Kai Tittmann
Reaction mechanisms of thiamin diphosphate enzymes: redox reactions.
FEBS J: 2009, 276(9);2454-68
[PubMed:19476487]
[WorldCat.org]
[DOI]
(I p)
U Neveling, S Bringer-Meyer, H Sahm
Gene and subunit organization of bacterial pyruvate dehydrogenase complexes.
Biochim Biophys Acta: 1998, 1385(2);367-72
[PubMed:9655937]
[WorldCat.org]
[DOI]
(P p)
M S Patel, T E Roche
Molecular biology and biochemistry of pyruvate dehydrogenase complexes.
FASEB J: 1990, 4(14);3224-33
[PubMed:2227213]
[WorldCat.org]
[DOI]
(P p)
P A Frey
Mechanism of coupled electron and group transfer in Escherichia coli pyruvate dehydrogenase.
Ann N Y Acad Sci: 1982, 378;250-64
[PubMed:6805383]
[WorldCat.org]
[DOI]
(P p)
Original publications
Leigh G Monahan, Isabella V Hajduk, Sinead P Blaber, Ian G Charles, Elizabeth J Harry
Coordinating bacterial cell division with nutrient availability: a role for glycolysis.
mBio: 2014, 5(3);e00935-14
[PubMed:24825009]
[WorldCat.org]
[DOI]
(I e)
Irnov Irnov, Cynthia M Sharma, Jörg Vogel, Wade C Winkler
Identification of regulatory RNAs in Bacillus subtilis.
Nucleic Acids Res: 2010, 38(19);6637-51
[PubMed:20525796]
[WorldCat.org]
[DOI]
(I p)
Shigeo Tojo, Kanako Kumamoto, Kazutake Hirooka, Yasutaro Fujita
Heavy involvement of stringent transcription control depending on the adenine or guanine species of the transcription initiation site in glucose and pyruvate metabolism in Bacillus subtilis.
J Bacteriol: 2010, 192(6);1573-85
[PubMed:20081037]
[WorldCat.org]
[DOI]
(I p)
Christine Eymann, Annette Dreisbach, Dirk Albrecht, Jörg Bernhardt, Dörte Becher, Sandy Gentner, Le Thi Tam, Knut Büttner, Gerrit Buurman, Christian Scharf, Simone Venz, Uwe Völker, Michael Hecker
A comprehensive proteome map of growing Bacillus subtilis cells.
Proteomics: 2004, 4(10);2849-76
[PubMed:15378759]
[WorldCat.org]
[DOI]
(P p)
Hans-Matti Blencke, Georg Homuth, Holger Ludwig, Ulrike Mäder, Michael Hecker, Jörg Stülke
Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways.
Metab Eng: 2003, 5(2);133-49
[PubMed:12850135]
[WorldCat.org]
[DOI]
(P p)
Haichun Gao, Xin Jiang, Kit Pogliano, Arthur I Aronson
The E1beta and E2 subunits of the Bacillus subtilis pyruvate dehydrogenase complex are involved in regulation of sporulation.
J Bacteriol: 2002, 184(10);2780-8
[PubMed:11976308]
[WorldCat.org]
[DOI]
(P p)
M M Nakano, Y P Dailly, P Zuber, D P Clark
Characterization of anaerobic fermentative growth of Bacillus subtilis: identification of fermentation end products and genes required for growth.
J Bacteriol: 1997, 179(21);6749-55
[PubMed:9352926]
[WorldCat.org]
[DOI]
(P p)
P N Lowe, J A Hodgson, R N Perham
Dual role of a single multienzyme complex in the oxidative decarboxylation of pyruvate and branched-chain 2-oxo acids in Bacillus subtilis.
Biochem J: 1983, 215(1);133-40
[PubMed:6414463]
[WorldCat.org]
[DOI]
(P p)