Difference between revisions of "PrpC"
| Line 15: | Line 15: | ||
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU15760 prpC] | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU15760 prpC] | ||
| + | |- | ||
| + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://subtiwiki.uni-goettingen.de/interact/ ''Subt''Interact]''': [http://subtiwiki.uni-goettingen.de/interact/index.php?protein=PrpC PrpC] | ||
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/subtipathways/search.php?enzyme=prpC prpC]''' | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/subtipathways/search.php?enzyme=prpC prpC]''' | ||
| Line 92: | Line 94: | ||
* '''[[SubtInteract|Interactions]]:''' | * '''[[SubtInteract|Interactions]]:''' | ||
| + | ** [[YvcK]]-[[PrpC]] {{PubMed|25012659}} | ||
* '''[[Localization]]:''' | * '''[[Localization]]:''' | ||
| Line 115: | Line 118: | ||
* '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=prpC_1650384_1651148_1 prpC] {{PubMed|22383849}} | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=prpC_1650384_1651148_1 prpC] {{PubMed|22383849}} | ||
| − | * '''Sigma factor:''' | + | * '''[[Sigma factor]]:''' |
* '''Regulation:''' | * '''Regulation:''' | ||
| Line 148: | Line 151: | ||
<pubmed> 21372323 </pubmed> | <pubmed> 21372323 </pubmed> | ||
==Original publications== | ==Original publications== | ||
| − | <pubmed>19246764,10986276,17693724 , 12399479, 18757537, 15530359 23793375 16025310 24390483</pubmed> | + | <pubmed>19246764,10986276,17693724 , 12399479, 18757537, 15530359 23793375 16025310 24390483 25012659</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 09:51, 12 July 2014
- Description: protein phosphatase
| Gene name | prpC |
| Synonyms | yloO |
| Essential | no |
| Product | protein phosphatase |
| Function | antagonist of PrkC-dependent phosphorylation |
| Gene expression levels in SubtiExpress: prpC | |
| Interactions involving this protein in SubtInteract: PrpC | |
| Metabolic function and regulation of this protein in SubtiPathways: prpC | |
| MW, pI | 27 kDa, 4.355 |
| Gene length, protein length | 762 bp, 254 aa |
| Immediate neighbours | yloN, prkC |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 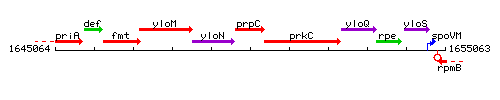
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed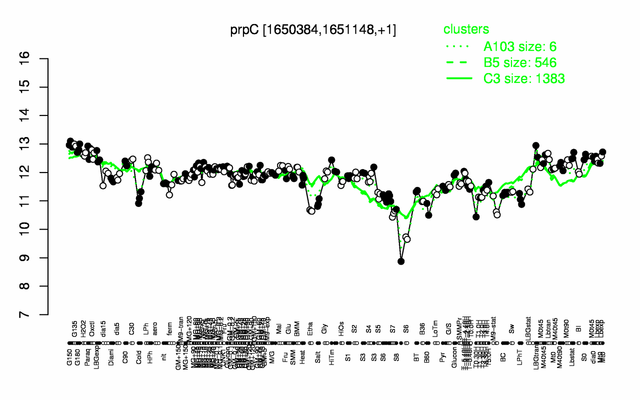
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU15760
Phenotypes of a mutant
A prpC mutant is less lytic in late stationary phase. PubMed
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: A phosphoprotein + H2O = a protein + phosphate (according to Swiss-Prot)
- Protein family:
- Paralogous protein(s):
Proteins dephosphorylated by PrpC
CpgA, EF-Tu, YezB PubMed, HPr PubMed, YkwC PubMed
Extended information on the protein
- Kinetic information:
- Modification:
- Cofactors: divalent cations such as magnesium or manganese
- Effectors of protein activity: inhibited by inorganic phosphate and glycero-2-phosphate PubMed
Database entries
- UniProt: O34779
- KEGG entry: [2]
- E.C. number: 3.1.3.16
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
- number of protein molecules per cell (complex medium with amino acids, without glucose): 459 PubMed
Biological materials
- Mutant: OMG401 (aphA3), available in the Stülke lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Sandro F F Pereira, Lindsie Goss, Jonathan Dworkin
Eukaryote-like serine/threonine kinases and phosphatases in bacteria.
Microbiol Mol Biol Rev: 2011, 75(1);192-212
[PubMed:21372323]
[WorldCat.org]
[DOI]
(I p)
Original publications