Difference between revisions of "YoaJ"
| Line 61: | Line 61: | ||
=== Additional information=== | === Additional information=== | ||
| − | |||
| − | |||
| − | |||
=The protein= | =The protein= | ||
| Line 79: | Line 76: | ||
* '''Kinetic information:''' | * '''Kinetic information:''' | ||
| − | * '''Domains:''' | + | * '''[[Domains]]:''' |
* '''Modification:''' | * '''Modification:''' | ||
| − | * ''' | + | * '''[[Cofactors]]:''' |
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| Line 140: | Line 137: | ||
=References= | =References= | ||
| − | + | <pubmed>19058186 15604683 16984999 22927418 18971341,21454649 12354229,23053073 24755657 22949138</pubmed> | |
| − | <pubmed>19058186 15604683 16984999 22927418 18971341,21454649 12354229,23053073</pubmed> | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 05:21, 25 April 2014
- Description: bacterial expansin, binds cellulose, required for the colonization of maize roots
| Gene name | yoaJ |
| Synonyms | |
| Essential | no |
| Product | expansin |
| Function | interaction with plant roots |
| Gene expression levels in SubtiExpress: yoaJ | |
| MW, pI | 25 kDa, 9.447 |
| Gene length, protein length | 696 bp, 232 aa |
| Immediate neighbours | yoaI, yoaK |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 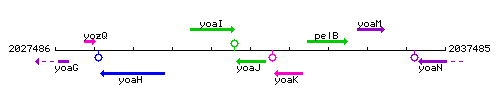
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed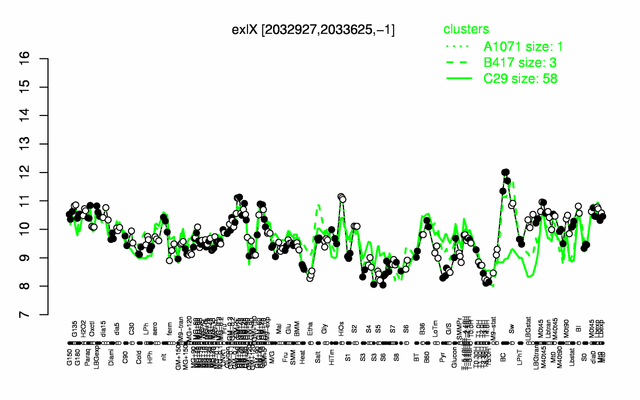
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU18630
Phenotypes of a mutant
strongly reduced ability to colonize maize roots PubMed
Database entries
- BsubCyc: BSU18630
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: expansin PubMed
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- Effectors of protein activity:
- Localization:
- extracellular PubMed
Database entries
- BsubCyc: BSU18630
- UniProt: O34918
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: yoaJ PubMed
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Miguel Olarte-Lozano, Mario A Mendoza-Nuñez, Nina Pastor, Lorenzo Segovia, Jorge Folch-Mallol, Claudia Martínez-Anaya
PcExl1 a novel acid expansin-like protein from the plant pathogen Pectobacterium carotovorum, binds cell walls differently to BsEXLX1.
PLoS One: 2014, 9(4);e95638
[PubMed:24755657]
[WorldCat.org]
[DOI]
(I e)
In Jung Kim, Hyeok-Jin Ko, Tae-Wan Kim, Ki Hyun Nam, In-Geol Choi, Kyoung Heon Kim
Binding characteristics of a bacterial expansin (BsEXLX1) for various types of pretreated lignocellulose.
Appl Microbiol Biotechnol: 2013, 97(12);5381-8
[PubMed:23053073]
[WorldCat.org]
[DOI]
(I p)
In Jung Kim, Hyeok-Jin Ko, Tae-Wan Kim, In-Geol Choi, Kyoung Heon Kim
Characteristics of the binding of a bacterial expansin (BsEXLX1) to microcrystalline cellulose.
Biotechnol Bioeng: 2013, 110(2);401-7
[PubMed:22949138]
[WorldCat.org]
[DOI]
(I p)
Nikolaos Georgelis, Neela H Yennawar, Daniel J Cosgrove
Structural basis for entropy-driven cellulose binding by a type-A cellulose-binding module (CBM) and bacterial expansin.
Proc Natl Acad Sci U S A: 2012, 109(37);14830-5
[PubMed:22927418]
[WorldCat.org]
[DOI]
(I p)
Nikolaos Georgelis, Akira Tabuchi, Nikolas Nikolaidis, Daniel J Cosgrove
Structure-function analysis of the bacterial expansin EXLX1.
J Biol Chem: 2011, 286(19);16814-23
[PubMed:21454649]
[WorldCat.org]
[DOI]
(I p)
Eun Sil Kim, Hee Jin Lee, Won-Gi Bang, In-Geol Choi, Kyoung Heon Kim
Functional characterization of a bacterial expansin from Bacillus subtilis for enhanced enzymatic hydrolysis of cellulose.
Biotechnol Bioeng: 2009, 102(5);1342-53
[PubMed:19058186]
[WorldCat.org]
[DOI]
(I p)
Frédéric Kerff, Ana Amoroso, Raphaël Herman, Eric Sauvage, Stéphanie Petrella, Patrice Filée, Paulette Charlier, Bernard Joris, Akira Tabuchi, Nikolas Nikolaidis, Daniel J Cosgrove
Crystal structure and activity of Bacillus subtilis YoaJ (EXLX1), a bacterial expansin that promotes root colonization.
Proc Natl Acad Sci U S A: 2008, 105(44);16876-81
[PubMed:18971341]
[WorldCat.org]
[DOI]
(I p)
Neela H Yennawar, Lian-Chao Li, David M Dudzinski, Akira Tabuchi, Daniel J Cosgrove
Crystal structure and activities of EXPB1 (Zea m 1), a beta-expansin and group-1 pollen allergen from maize.
Proc Natl Acad Sci U S A: 2006, 103(40);14664-71
[PubMed:16984999]
[WorldCat.org]
[DOI]
(P p)
Hans Kende, Kent Bradford, David Brummell, Hyung-Taeg Cho, Daniel Cosgrove, Andrew Fleming, Chris Gehring, Yi Lee, Simon McQueen-Mason, Jocelyn Rose, Laurentius A C J Voesenek
Nomenclature for members of the expansin superfamily of genes and proteins.
Plant Mol Biol: 2004, 55(3);311-4
[PubMed:15604683]
[WorldCat.org]
[DOI]
(P p)
Noel Baichoo, Tao Wang, Rick Ye, John D Helmann
Global analysis of the Bacillus subtilis Fur regulon and the iron starvation stimulon.
Mol Microbiol: 2002, 45(6);1613-29
[PubMed:12354229]
[WorldCat.org]
[DOI]
(P p)