Difference between revisions of "PbpE"
| Line 40: | Line 40: | ||
{{SubtiWiki category|[[cell wall synthesis]]}}, | {{SubtiWiki category|[[cell wall synthesis]]}}, | ||
{{SubtiWiki category|[[cell wall degradation/ turnover]]}}, | {{SubtiWiki category|[[cell wall degradation/ turnover]]}}, | ||
| − | {{SubtiWiki category|[[cell envelope stress proteins (controlled by SigM, V, W, X, Y)]]}} | + | {{SubtiWiki category|[[cell envelope stress proteins (controlled by SigM, V, W, X, Y)]]}}, |
| + | [[most abundant proteins]] | ||
= This gene is a member of the following [[regulons]] = | = This gene is a member of the following [[regulons]] = | ||
| Line 60: | Line 61: | ||
=== Additional information=== | === Additional information=== | ||
| − | |||
| − | |||
| − | |||
=The protein= | =The protein= | ||
| Line 78: | Line 76: | ||
* '''Kinetic information:''' | * '''Kinetic information:''' | ||
| − | * '''Domains:''' | + | * '''[[Domains]]:''' |
* '''Modification:''' | * '''Modification:''' | ||
| − | * ''' | + | * '''[[Cofactors]]:''' |
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| Line 115: | Line 113: | ||
* '''Regulatory mechanism:''' | * '''Regulatory mechanism:''' | ||
| − | * '''Additional information:''' the mRNA is very stable (half-life > 15 min) [http://www.ncbi.nlm.nih.gov/sites/entrez/12884008 PubMed] | + | * '''Additional information:''' |
| + | ** the mRNA is very stable (half-life > 15 min) [http://www.ncbi.nlm.nih.gov/sites/entrez/12884008 PubMed] | ||
| + | ** belongs to the 100 [[most abundant proteins]] {{PubMed|15378759}} | ||
=Biological materials = | =Biological materials = | ||
| Line 137: | Line 137: | ||
=References= | =References= | ||
| − | <pubmed>8491712,12533473,9987136,19063962,12884008,12207695,7592498, 20817675</pubmed> | + | <pubmed>8491712,12533473,9987136,19063962,12884008,12207695,7592498, 20817675 15378759</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 16:32, 5 March 2014
- Description: penicillin-binding protein PBP 4* (spore cortex)
| Gene name | pbpE |
| Synonyms | |
| Essential | no |
| Product | penicillin-binding protein PBP 4* (spore cortex) |
| Function | endopeptidase |
| Gene expression levels in SubtiExpress: pbpE | |
| MW, pI | 51 kDa, 4.835 |
| Gene length, protein length | 1353 bp, 451 aa |
| Immediate neighbours | racX, sacB |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 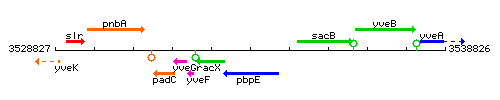
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed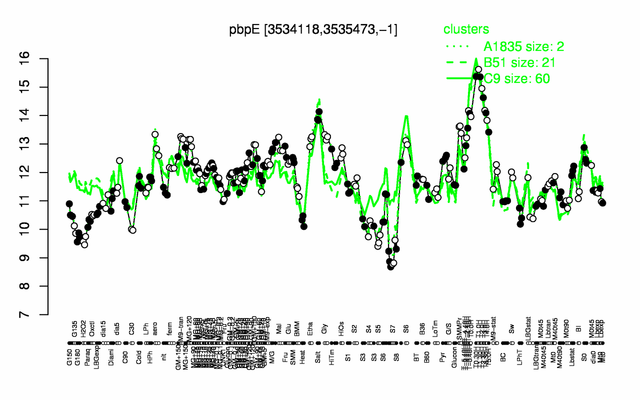
| |
Contents
Categories containing this gene/protein
cell wall synthesis, cell wall degradation/ turnover, cell envelope stress proteins (controlled by SigM, V, W, X, Y), most abundant proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU34440
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: beta-lactamase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- Effectors of protein activity:
- Localization: spore cortex
Database entries
- Structure: 3TG9 (from Bacillus halodurans, 27% identity, 59% similarity)
- UniProt: P32959
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
- the mRNA is very stable (half-life > 15 min) PubMed
- belongs to the 100 most abundant proteins PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Onuma Chumsakul, Hiroki Takahashi, Taku Oshima, Takahiro Hishimoto, Shigehiko Kanaya, Naotake Ogasawara, Shu Ishikawa
Genome-wide binding profiles of the Bacillus subtilis transition state regulator AbrB and its homolog Abh reveals their interactive role in transcriptional regulation.
Nucleic Acids Res: 2011, 39(2);414-28
[PubMed:20817675]
[WorldCat.org]
[DOI]
(I p)
María Mercedes Palomino, Carmen Sanchez-Rivas, Sandra M Ruzal
High salt stress in Bacillus subtilis: involvement of PBP4* as a peptidoglycan hydrolase.
Res Microbiol: 2009, 160(2);117-24
[PubMed:19063962]
[WorldCat.org]
[DOI]
(P p)
Christine Eymann, Annette Dreisbach, Dirk Albrecht, Jörg Bernhardt, Dörte Becher, Sandy Gentner, Le Thi Tam, Knut Büttner, Gerrit Buurman, Christian Scharf, Simone Venz, Uwe Völker, Michael Hecker
A comprehensive proteome map of growing Bacillus subtilis cells.
Proteomics: 2004, 4(10);2849-76
[PubMed:15378759]
[WorldCat.org]
[DOI]
(P p)
G Hambraeus, C von Wachenfeldt, L Hederstedt
Genome-wide survey of mRNA half-lives in Bacillus subtilis identifies extremely stable mRNAs.
Mol Genet Genomics: 2003, 269(5);706-14
[PubMed:12884008]
[WorldCat.org]
[DOI]
(P p)
Stephan Zellmeier, Ulrich Zuber, Wolfgang Schumann, Thomas Wiegert
The absence of FtsH metalloprotease activity causes overexpression of the sigmaW-controlled pbpE gene, resulting in filamentous growth of Bacillus subtilis.
J Bacteriol: 2003, 185(3);973-82
[PubMed:12533473]
[WorldCat.org]
[DOI]
(P p)
Min Cao, Tao Wang, Rick Ye, John D Helmann
Antibiotics that inhibit cell wall biosynthesis induce expression of the Bacillus subtilis sigma(W) and sigma(M) regulons.
Mol Microbiol: 2002, 45(5);1267-76
[PubMed:12207695]
[WorldCat.org]
[DOI]
(P p)
X Huang, A Gaballa, M Cao, J D Helmann
Identification of target promoters for the Bacillus subtilis extracytoplasmic function sigma factor, sigma W.
Mol Microbiol: 1999, 31(1);361-71
[PubMed:9987136]
[WorldCat.org]
[DOI]
(P p)
M A Strauch
Delineation of AbrB-binding sites on the Bacillus subtilis spo0H, kinB, ftsAZ, and pbpE promoters and use of a derived homology to identify a previously unsuspected binding site in the bsuB1 methylase promote.
J Bacteriol: 1995, 177(23);6999-7002
[PubMed:7592498]
[WorldCat.org]
[DOI]
(P p)
D L Popham, P Setlow
Cloning, nucleotide sequence, and regulation of the Bacillus subtilis pbpE operon, which codes for penicillin-binding protein 4* and an apparent amino acid racemase.
J Bacteriol: 1993, 175(10);2917-25
[PubMed:8491712]
[WorldCat.org]
[DOI]
(P p)