Difference between revisions of "DacB"
| Line 123: | Line 123: | ||
* '''Mutant:''' | * '''Mutant:''' | ||
| + | ** 1A744 ( ''dacB''::''cat''), {{PubMed|1644769}}, available at [http://pasture.asc.ohio-state.edu/BGSC/getdetail.cfm?bgscid=1A744&Search=1A744 BGSC] | ||
* '''Expression vector:''' | * '''Expression vector:''' | ||
Revision as of 09:04, 19 September 2012
- Description: sporulation-specific penicillin-binding protein 5, D-alanyl-D-alanine carboxypeptidase
| Gene name | dacB |
| Synonyms | |
| Essential | no |
| Product | penicillin-binding protein 5, D-alanyl-D-alanine carboxypeptidase |
| Function | carboxypeptidase |
| Gene expression levels in SubtiExpress: dacB | |
| MW, pI | 42 kDa, 9.725 |
| Gene length, protein length | 1146 bp, 382 aa |
| Immediate neighbours | spmA, ypuI |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 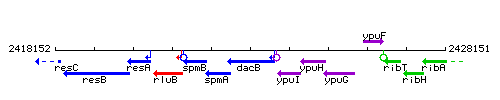
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed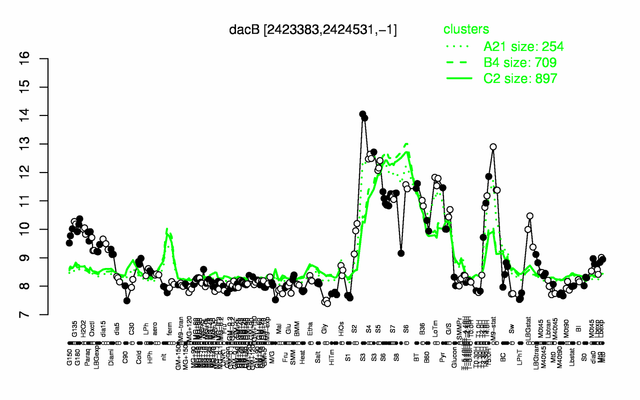
| |
Contents
Categories containing this gene/protein
cell wall synthesis, sporulation proteins, membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU23190
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: modifies degree of cross-linking of glycan strands in peptidoglycan
- Protein family: peptidase S11 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- cell membrane (according to Swiss-Prot)
Database entries
- Structure: 3MFD (beta-lactamase superfamily domain)
- UniProt: P35150
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulatory mechanism:
- Additional information:
Biological materials
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Additional publications: PubMed