Difference between revisions of "YxxG"
(→Extended information on the protein) |
|||
| Line 100: | Line 100: | ||
* '''Operon:''' ''[[wapA]]-[[yxxG]]'' [http://www.ncbi.nlm.nih.gov/sites/entrez/16306698 PubMed] | * '''Operon:''' ''[[wapA]]-[[yxxG]]'' [http://www.ncbi.nlm.nih.gov/sites/entrez/16306698 PubMed] | ||
| − | * '''[ | + | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=yxxG_4023054_4023482_-1 yxxG] {{PubMed|22383849}} |
| + | |||
| + | * '''Sigma factor:''' | ||
* '''Regulation:''' | * '''Regulation:''' | ||
Revision as of 09:49, 17 April 2012
- Description: unknown
| Gene name | yxxG |
| Synonyms | |
| Essential | no |
| Product | unknown |
| Function | unknown |
| MW, pI | 16 kDa, 4.424 |
| Gene length, protein length | 426 bp, 142 aa |
| Immediate neighbours | yxiF, wapA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 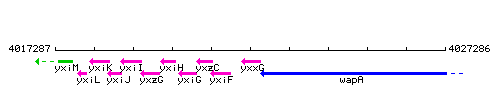
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
phosphoproteins, proteins of unknown function
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU39220
Phenotypes of a mutant
- severe growth defect PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification: phosphorylation on (Tyr-102 OR Ser-105) PubMed
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: Q07836
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Sigma factor:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Taryn B Kiley, Nicola R Stanley-Wall
Post-translational control of Bacillus subtilis biofilm formation mediated by tyrosine phosphorylation.
Mol Microbiol: 2010, 78(4);947-63
[PubMed:20815827]
[WorldCat.org]
[DOI]
(I p)
Boris Macek, Ivan Mijakovic, Jesper V Olsen, Florian Gnad, Chanchal Kumar, Peter R Jensen, Matthias Mann
The serine/threonine/tyrosine phosphoproteome of the model bacterium Bacillus subtilis.
Mol Cell Proteomics: 2007, 6(4);697-707
[PubMed:17218307]
[WorldCat.org]
[DOI]
(P p)
Juliane Ollinger, Kyung-Bok Song, Haike Antelmann, Michael Hecker, John D Helmann
Role of the Fur regulon in iron transport in Bacillus subtilis.
J Bacteriol: 2006, 188(10);3664-73
[PubMed:16672620]
[WorldCat.org]
[DOI]
(P p)
Masakuni Serizawa, Keisuke Kodama, Hiroki Yamamoto, Kazuo Kobayashi, Naotake Ogasawara, Junichi Sekiguchi
Functional analysis of the YvrGHb two-component system of Bacillus subtilis: identification of the regulated genes by DNA microarray and northern blot analyses.
Biosci Biotechnol Biochem: 2005, 69(11);2155-69
[PubMed:16306698]
[WorldCat.org]
[DOI]
(P p)
V Dartois, M Débarbouillé, F Kunst, G Rapoport
Characterization of a novel member of the DegS-DegU regulon affected by salt stress in Bacillus subtilis.
J Bacteriol: 1998, 180(7);1855-61
[PubMed:9537385]
[WorldCat.org]
[DOI]
(P p)