Difference between revisions of "CymR"
(→Categories containing this gene/protein) |
|||
| Line 1: | Line 1: | ||
| − | * '''Description:''' pleiotropic regulator of | + | * '''Description:''' pleiotropic regulator of [[sulfur metabolism]] <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
| Line 12: | Line 12: | ||
|style="background:#ABCDEF;" align="center"| '''Product''' || transcription repressor | |style="background:#ABCDEF;" align="center"| '''Product''' || transcription repressor | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Function''' || regulation of | + | |style="background:#ABCDEF;" align="center"|'''Function''' || regulation of [[sulfur metabolism]] |
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/riboflavin.html Riboflavin / FAD], [http://subtiwiki.uni-goettingen.de/pathways/cys_meth_and_sulfate_assimilation.html Cys, Met & Sulfate assimilation]''' | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/riboflavin.html Riboflavin / FAD], [http://subtiwiki.uni-goettingen.de/pathways/cys_meth_and_sulfate_assimilation.html Cys, Met & Sulfate assimilation]''' | ||
| Line 59: | Line 59: | ||
=== Additional information=== | === Additional information=== | ||
| − | |||
| − | |||
| Line 88: | Line 86: | ||
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| − | * '''Interactions:''' [[CymR]]-[[CysK]], the complex is formed in the presence of cysteine, and results in DNA binding and repression of genes of cysteine biosynthesis [http://www.ncbi.nlm.nih.gov/sites/entrez/18974048 PubMed] | + | * '''Interactions:''' |
| + | ** [[CymR]]-[[CysK]], the complex is formed in the presence of cysteine, and results in DNA binding and repression of genes of cysteine biosynthesis [http://www.ncbi.nlm.nih.gov/sites/entrez/18974048 PubMed] | ||
* '''Localization:''' cytoplasm (according to Swiss-Prot) | * '''Localization:''' cytoplasm (according to Swiss-Prot) | ||
| Line 94: | Line 93: | ||
=== Database entries === | === Database entries === | ||
| − | * '''Structure:''' | + | * '''Structure:''' {{PubMed|21624051}} |
* '''UniProt:''' [http://www.uniprot.org/uniprot/O34527 O34527] | * '''UniProt:''' [http://www.uniprot.org/uniprot/O34527 O34527] | ||
| Line 140: | Line 139: | ||
<pubmed>16513748, </pubmed> | <pubmed>16513748, </pubmed> | ||
==Other original publications== | ==Other original publications== | ||
| − | <pubmed>18974048,17056751,16109943,16885442 11948165 , 15668000 20608979 </pubmed> | + | <pubmed>18974048,17056751,16109943,16885442 11948165 , 15668000 20608979 21624051</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 08:35, 2 June 2011
- Description: pleiotropic regulator of sulfur metabolism
| Gene name | cymR |
| Synonyms | yrzC |
| Essential | no |
| Product | transcription repressor |
| Function | regulation of sulfur metabolism |
| Metabolic function and regulation of this protein in SubtiPathways: Riboflavin / FAD, Cys, Met & Sulfate assimilation | |
| MW, pI | 11 kDa, 9.879 |
| Gene length, protein length | 333 bp, 111 aa |
| Immediate neighbours | yrvO, yrvN |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 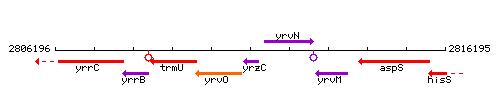
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
sulfur metabolism, transcription factors and their control, regulators of core metabolism
This gene is a member of the following regulons
The CymR regulon
The gene
Basic information
- Locus tag: BSU27520
Phenotypes of a mutant
- impaired growth in the presence of cystine and increased sensitivity to hydrogen peroxide-, disulfide-, paraquat- and tellurite-induced stresses PubMed
- increased intracellular cysteine pool and hydrogen sulfide formation PubMed
- depletion of branched-chain amino acids PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: Rrf2 family of transcription regulators
- Paralogous protein(s):
Genes repressed by CymR
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Effectors of protein activity:
- Interactions:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- Structure: PubMed
- UniProt: O34527
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Isabelle Martin-Verstraete, Institute Pasteur, Paris, France
Your additional remarks
References
The CymR regulon
Other original publications