Difference between revisions of "DivIB"
(→References) |
(→References) |
||
| Line 142: | Line 142: | ||
=References= | =References= | ||
| − | <pubmed>8459777,10852898, 19429628 1391053 21219466 8320223 8106328 18179421 | + | '''Additional references:''' {{PubMed|20870765}} |
| − | + | <pubmed>8459777,10852898, 19429628 1391053 21219466 8320223 8106328 18179421 15659156,16936019,10217758, 10792716,9720875, 16936026,9426141, 9044263,</pubmed> | |
| − | + | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 10:20, 12 January 2011
- Description: cell-division initiation protein (septum formation)
| Gene name | divIB |
| Synonyms | dds, tms-12 |
| Essential | yes PubMed |
| Product | cell-division protein |
| Function | septum formation |
| Metabolic function and regulation of this protein in SubtiPathways: Cell wall | |
| MW, pI | 30 kDa, 9.244 |
| Gene length, protein length | 789 bp, 263 aa |
| Immediate neighbours | murB, ylxW |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 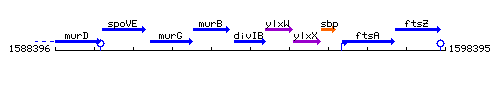
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
cell division, sporulation proteins, cell envelope stress proteins (controlled by SigM, W, X, Y), essential genes, membrane proteins
This gene is a member of the following regulons
SigE regulon, SigM regulon, SpoIIID regulon
The gene
Basic information
- Locus tag: BSU15240
Phenotypes of a mutant
essential PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: ftsQ family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: cell membrane PubMed
Database entries
- Structure: 2ALJ (major extracytoplasmic domain, Geobacillus stearothermophilus)
- UniProt: P16655
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Additional references: PubMed