Difference between revisions of "XynA"
(→References) |
(→References) |
||
| Line 121: | Line 121: | ||
=References= | =References= | ||
| − | <pubmed>19531602 ,8012596, 18957862, 19422059 19994888 </pubmed> | + | <pubmed>19531602 ,8012596, 18957862, 19422059 19994888 20096384 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 10:24, 26 January 2010
- Description: endo-1,4-beta-xylanase
| Gene name | xynA |
| Synonyms | |
| Essential | no |
| Product | endo-1,4-beta-xylanase |
| Function | xylan degradation |
| Metabolic function and regulation of this protein in SubtiPathways: Sugar catabolism | |
| MW, pI | 23 kDa, 9.644 |
| Gene length, protein length | 639 bp, 213 aa |
| Immediate neighbours | pps, yobD |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 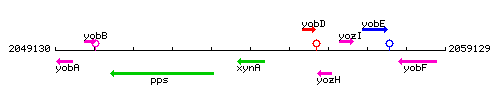
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU18840
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Endohydrolysis of (1->4)-beta-D-xylosidic linkages in xylans (according to Swiss-Prot)
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: extracellular (signal peptide) PubMed
Database entries
- UniProt: P18429
- KEGG entry: [3]
- E.C. number: 3.2.1.8
Additional information
Expression and regulation
- Operon: xynA PubMed
- Regulation:
- constitutively expressed PubMed
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Annick Pollet, Stijn Lagaert, Elena Eneyskaya, Anna Kulminskaya, Jan A Delcour, Christophe M Courtin
Mutagenesis and subsite mapping underpin the importance for substrate specificity of the aglycon subsites of glycoside hydrolase family 11 xylanases.
Biochim Biophys Acta: 2010, 1804(4);977-85
[PubMed:20096384]
[WorldCat.org]
[DOI]
(P p)
Louise E Rasmussen, Anne S Meyer
Size exclusion chromatography for the quantitative profiling of the enzyme-catalyzed hydrolysis of Xylo-oligosaccharides.
J Agric Food Chem: 2010, 58(2);762-9
[PubMed:19994888]
[WorldCat.org]
[DOI]
(I p)
Tim Beliën, Iris J Joye, Jan A Delcour, Christophe M Courtin
Computational design-based molecular engineering of the glycosyl hydrolase family 11 B. subtilis XynA endoxylanase improves its acid stability.
Protein Eng Des Sel: 2009, 22(10);587-96
[PubMed:19531602]
[WorldCat.org]
[DOI]
(I p)
Annick Pollet, Elien Vandermarliere, Jeroen Lammertyn, Sergei V Strelkov, Jan A Delcour, Christophe M Courtin
Crystallographic and activity-based evidence for thumb flexibility and its relevance in glycoside hydrolase family 11 xylanases.
Proteins: 2009, 77(2);395-403
[PubMed:19422059]
[WorldCat.org]
[DOI]
(I p)
Birgit Voigt, Haike Antelmann, Dirk Albrecht, Armin Ehrenreich, Karl-Heinz Maurer, Stefan Evers, Gerhard Gottschalk, Jan Maarten van Dijl, Thomas Schweder, Michael Hecker
Cell physiology and protein secretion of Bacillus licheniformis compared to Bacillus subtilis.
J Mol Microbiol Biotechnol: 2009, 16(1-2);53-68
[PubMed:18957862]
[WorldCat.org]
[DOI]
(I p)
C Lindner, J Stülke, M Hecker
Regulation of xylanolytic enzymes in Bacillus subtilis.
Microbiology (Reading): 1994, 140 ( Pt 4);753-7
[PubMed:8012596]
[WorldCat.org]
[DOI]
(P p)