Difference between revisions of "YjmC"
| Line 1: | Line 1: | ||
| − | * '''Description:''' | + | * '''Description:''' unknown, may be involved in galacturonate utilization <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
| Line 85: | Line 85: | ||
=== Additional information=== | === Additional information=== | ||
| + | The gene is annotated in KEGG as an ortholog of malate dehydrogenase EC 1.1.1.37. It is marked but is marked as “uncharacterized oxidoreductase” (EC 1.1.1.-) in Swiss-Prot. In MetaCyc it is marked as “similar to malate dehydrogenase”. As the paper by Mekjian et al. {{PubMed|9882655}} suggests this gene is more likely to be involved in the glucuronate pathway (for which EC 1.1.1.37 is not a member), no literature/experimental evidence supporting the annotation is available. {{PubMed|19935659}} | ||
=Expression and regulation= | =Expression and regulation= | ||
Revision as of 12:01, 26 November 2009
- Description: unknown, may be involved in galacturonate utilization
| Gene name | yjmC |
| Synonyms | |
| Essential | no |
| Product | unknown |
| Function | unknown |
| MW, pI | 36 kDa, 5.669 |
| Gene length, protein length | 1011 bp, 337 aa |
| Immediate neighbours | yjmB, yjmD |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 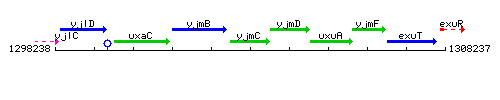
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU12320
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: LDH2/MDH2 oxidoreductase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- Structure:
- UniProt: O34736
- KEGG entry: [3]
- E.C. number:
Additional information
The gene is annotated in KEGG as an ortholog of malate dehydrogenase EC 1.1.1.37. It is marked but is marked as “uncharacterized oxidoreductase” (EC 1.1.1.-) in Swiss-Prot. In MetaCyc it is marked as “similar to malate dehydrogenase”. As the paper by Mekjian et al. PubMed suggests this gene is more likely to be involved in the glucuronate pathway (for which EC 1.1.1.37 is not a member), no literature/experimental evidence supporting the annotation is available. PubMed
Expression and regulation
- Operon:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References