Difference between revisions of "FlhG"
(→References) |
|||
| (48 intermediate revisions by 6 users not shown) | |||
| Line 1: | Line 1: | ||
| − | * '''Description:''' | + | * '''Description:''' GTPase activating protein, activates [[FlhF]], activates assembly of the flagellar C ring, control of flagellar basal body position <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Gene name''' | |style="background:#ABCDEF;" align="center"|'''Gene name''' | ||
| − | |'' | + | |''flhG '' |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Synonyms''' || ''ylxH'' | + | |style="background:#ABCDEF;" align="center"| '''Synonyms''' || ''ylxH '' |
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''Essential''' || no | |style="background:#ABCDEF;" align="center"| '''Essential''' || no | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Product''' || | + | |style="background:#ABCDEF;" align="center"| '''Product''' || GTPase activating protein |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Function''' || | + | |style="background:#ABCDEF;" align="center"|'''Function''' || activation of [[FlhF]] |
| + | |- | ||
| + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU16410 flhG] | ||
| + | |- | ||
| + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://subtiwiki.uni-goettingen.de/interact/ ''Subt''Interact]''': [http://subtiwiki.uni-goettingen.de/interact/index.php?protein=FlhG FlhG] | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 33 kDa, 9.648 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 33 kDa, 9.648 | ||
| Line 20: | Line 24: | ||
|style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[flhF]]'', ''[[cheB]]'' | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[flhF]]'', ''[[cheB]]'' | ||
|- | |- | ||
| − | |style="background:#FAF8CC;" align="center"|'''[http:// | + | |style="background:#FAF8CC;" align="center"|'''Sequences'''||[http://bsubcyc.org/BSUB/sequence-aa?type=GENE&object=BSU16410 Protein] [http://bsubcyc.org/BSUB/sequence?type=GENE&object=BSU16410 DNA] [http://bsubcyc.org/BSUB/seq-selector?chromosome=CHROM-1&object=BSU16410 DNA_with_flanks] |
| − | |||
|- | |- | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:ylxH_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:ylxH_context.gif]] | ||
| + | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
| + | |- | ||
| + | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=ylxH_1710838_1711734_1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:ylxH_expression.png|500px|link=http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU16410]] | ||
|- | |- | ||
|} | |} | ||
__TOC__ | __TOC__ | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/> | ||
| − | + | = [[Categories]] containing this gene/protein = | |
| + | {{SubtiWiki category|[[motility and chemotaxis]]}}, | ||
| + | {{SubtiWiki category|[[resistance against toxins/ antibiotics]]}} | ||
| + | |||
| + | = This gene is a member of the following [[regulons]] = | ||
| + | {{SubtiWiki regulon|[[CodY regulon]]}}, | ||
| + | {{SubtiWiki regulon|[[DegU regulon]]}}, | ||
| + | {{SubtiWiki regulon|[[SigD regulon]]}}, | ||
| + | {{SubtiWiki regulon|[[Spo0A regulon]]}} | ||
=The gene= | =The gene= | ||
| Line 35: | Line 53: | ||
=== Basic information === | === Basic information === | ||
| − | * ''' | + | * '''Locus tag:''' BSU16410 |
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
| − | susceptible to acriflavine and ethidium bromide, and severe growth inhibition as surfactin concentration increased up to 100ug/ml. | + | * susceptible to acriflavine and ethidium bromide, and severe growth inhibition as surfactin concentration increased up to 100ug/ml. |
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU16410&redirect=T BSU16410] | ||
* '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/fla-che.html] | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/fla-che.html] | ||
| Line 55: | Line 74: | ||
* '''Catalyzed reaction/ biological activity:''' | * '''Catalyzed reaction/ biological activity:''' | ||
| + | ** binds [[FlhF]] and stimulates its GTPase activity {{PubMed|22056770}} | ||
| + | ** controls the distance between the flagellar bodies {{PubMed|23190039}} | ||
| − | * '''Protein family:''' | + | * '''Protein family:''' View classification (according to Swiss-Prot) |
* '''Paralogous protein(s):''' | * '''Paralogous protein(s):''' | ||
| Line 64: | Line 85: | ||
* '''Kinetic information:''' | * '''Kinetic information:''' | ||
| − | * '''Domains:''' | + | * '''[[Domains]]:''' |
* '''Modification:''' | * '''Modification:''' | ||
| Line 72: | Line 93: | ||
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| − | * '''Interactions:''' | + | * '''[[SubtInteract|Interactions]]:''' |
| + | ** forms homodimers {{PubMed|25733861}} | ||
| + | ** [[FlhG]]-[[FlhF]] ([[FlhG]] binds the GTP-bound homodimer of [[FlhF]]) {{PubMed|22056770}} | ||
| + | ** [[FlhG]]-([[FliY]]-[[FliM]])-[[FliG]] {{PubMed|25733861}} | ||
| − | * '''Localization:''' | + | * '''[[Localization]]:''' cytoplasm (homogeneous) and flagellar basal body {{PubMed|25733861,16479537}} |
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU16410&redirect=T BSU16410] | ||
| − | * '''Structure:''' | + | * '''Structure:''' |
| + | ** [http://www.rcsb.org/pdb/explore/explore.do?structureId=4RZ3 4RZ3] (from ''Geobacillus thermodenitrificans'') {{PubMed|25733861}} | ||
| + | ** [http://www.rcsb.org/pdb/explore.do?structureId=3SYN 3SYN] (complex with [[FlhF]]) {{PubMed|22056770}} | ||
| − | * ''' | + | * '''UniProt:''' |
| + | ** [http://www.uniprot.org/uniprot/P40742 P40742] | ||
| + | ** [http://www.rcsb.org/pdb/explore.do?structureId=3SYN 3SYN] (complex with [[FlhF]]) {{PubMed|22056770}} | ||
| − | * '''KEGG entry:''' | + | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu:BSU16410] |
* '''E.C. number:''' | * '''E.C. number:''' | ||
| Line 89: | Line 118: | ||
=Expression and regulation= | =Expression and regulation= | ||
| + | * '''Operon:''' part of the [[fla-che operon]] | ||
| + | ** ''[[flgB]]-[[flgC]]-[[fliE]]-[[fliF]]-[[fliG]]-[[fliH]]-[[fliI]]-[[fliJ]]-[[ylxF]]-[[fliK]]-[[flgD]]-[[flgE]]-[[fliL]]-[[fliM]]-[[fliY]]-[[cheY]]-[[fliZ]]-[[fliP]]-[[fliQ]]-[[fliR]]-[[flhB]]-[[flhA]]-[[flhF]]-[[flhG]]-[[cheB]]-[[cheA]]-[[cheW]]-[[cheC]]-[[cheD]]-[[sigD]]-[[swrB]]'' {{PubMed|9657996,8157612,15175317}} | ||
| + | ** ''[[ylxF]]-[[fliK]]-[[flgD]]-[[flgE]]-[[fliL]]-[[fliM]]-[[fliY]]-[[cheY]]-[[fliZ]]-[[fliP]]-[[fliQ]]-[[fliR]]-[[flhB]]-[[flhA]]-[[flhF]]-[[flhG]]-[[cheB]]-[[cheA]]-[[cheW]]-[[cheC]]-[[cheD]]-[[sigD]]-[[swrB]]'' {{PubMed|20233303}} | ||
| − | * ''' | + | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=ylxH_1710838_1711734_1 ylxH] {{PubMed|22383849}} |
| − | * '''Sigma factor:''' | + | * '''[[Sigma factor]]:''' |
| + | ** ''[[flgB]]'': [[SigA]] {{PubMed|9657996}}, [[SigD]] {{PubMed|9657996}} | ||
| + | ** ''[[ylxF]]'': [[SigD]] {{PubMed|20233303}} | ||
| − | * '''Regulation:''' | + | * '''Regulation:''' see [[fla-che operon]] |
| − | * '''Regulatory mechanism:''' | + | * '''Regulatory mechanism:''' see [[fla-che operon]] |
| − | * '''Additional information:''' | + | * '''Additional information:''' |
=Biological materials = | =Biological materials = | ||
| Line 119: | Line 153: | ||
=References= | =References= | ||
| + | ==Reviews== | ||
| + | <pubmed> 26195616,, </pubmed> | ||
| + | ==Original Publications== | ||
| + | <pubmed>15066026,17850253,14651647, 16479537 9657996,8157612,15175317 22056770 23190039 24386445 25733861</pubmed> | ||
| − | + | [[Category:Protein-coding genes]] | |
Latest revision as of 10:54, 24 July 2015
- Description: GTPase activating protein, activates FlhF, activates assembly of the flagellar C ring, control of flagellar basal body position
| Gene name | flhG |
| Synonyms | ylxH |
| Essential | no |
| Product | GTPase activating protein |
| Function | activation of FlhF |
| Gene expression levels in SubtiExpress: flhG | |
| Interactions involving this protein in SubtInteract: FlhG | |
| MW, pI | 33 kDa, 9.648 |
| Gene length, protein length | 894 bp, 298 aa |
| Immediate neighbours | flhF, cheB |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 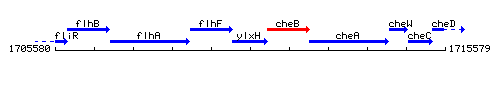
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed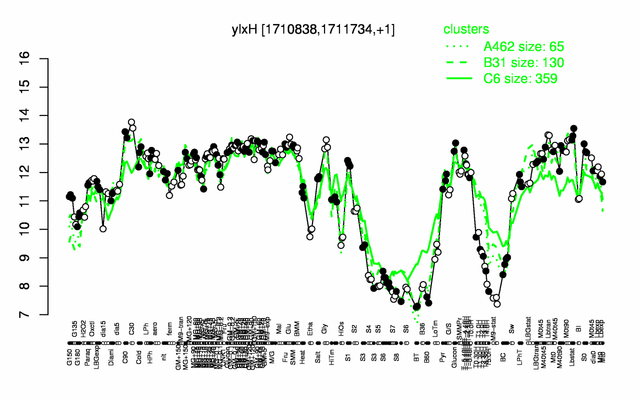
| |
Contents
Categories containing this gene/protein
motility and chemotaxis, resistance against toxins/ antibiotics
This gene is a member of the following regulons
CodY regulon, DegU regulon, SigD regulon, Spo0A regulon
The gene
Basic information
- Locus tag: BSU16410
Phenotypes of a mutant
- susceptible to acriflavine and ethidium bromide, and severe growth inhibition as surfactin concentration increased up to 100ug/ml.
Database entries
- BsubCyc: BSU16410
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: View classification (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: cytoplasm (homogeneous) and flagellar basal body PubMed
Database entries
- BsubCyc: BSU16410
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: part of the fla-che operon
- Regulation: see fla-che operon
- Regulatory mechanism: see fla-che operon
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original Publications