Difference between revisions of "OhrR"
(→References) |
|||
| (6 intermediate revisions by 3 users not shown) | |||
| Line 15: | Line 15: | ||
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU13150 ohrR] | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU13150 ohrR] | ||
| + | |- | ||
| + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://subtiwiki.uni-goettingen.de/interact/ ''Subt''Interact]''': [http://subtiwiki.uni-goettingen.de/interact/index.php?protein=OhrR OhrR] | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 16 kDa, 6.364 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 16 kDa, 6.364 | ||
| Line 35: | Line 37: | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
| − | |||
| − | |||
| − | |||
| − | |||
<br/><br/><br/> | <br/><br/><br/> | ||
| Line 58: | Line 56: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU13150&redirect=T BSU13150] | ||
* '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/ykmA.html] | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/ykmA.html] | ||
| Line 80: | Line 79: | ||
* '''Kinetic information:''' | * '''Kinetic information:''' | ||
| − | * '''Domains:''' | + | * '''[[Domains]]:''' |
* '''Modification:''' | * '''Modification:''' | ||
| Line 86: | Line 85: | ||
** S-bacillithiolated by NaOCl and CHP stress on Cys-15, this results in release from the'' [[ohrA]]'' promoter {{PubMed|21749987}} | ** S-bacillithiolated by NaOCl and CHP stress on Cys-15, this results in release from the'' [[ohrA]]'' promoter {{PubMed|21749987}} | ||
| − | * ''' | + | * '''[[Cofactors]]:''' |
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| Line 92: | Line 91: | ||
* '''[[SubtInteract|Interactions]]:''' | * '''[[SubtInteract|Interactions]]:''' | ||
** forms dimers | ** forms dimers | ||
| + | ** [[BrxA]]-[[OhrR]], to de-bacillithiolate S-bacillithiolated [[OhrR]] {{PubMed|24313874}} | ||
| + | ** [[BrxB]]-[[OhrR]], to de-bacillithiolate S-bacillithiolated [[OhrR]] {{PubMed|24313874}} | ||
* '''[[Localization]]:''' cytoplasm (according to Swiss-Prot) | * '''[[Localization]]:''' cytoplasm (according to Swiss-Prot) | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU13150&redirect=T BSU13150] | ||
* '''Structure:''' [http://www.rcsb.org/pdb/explore.do?structureId=1Z91 1Z91] (reduced form), [http://www.rcsb.org/pdb/explore.do?structureId=1Z9C 1Z9C] (complex with ''[[ohrA]]'' promoter) | * '''Structure:''' [http://www.rcsb.org/pdb/explore.do?structureId=1Z91 1Z91] (reduced form), [http://www.rcsb.org/pdb/explore.do?structureId=1Z9C 1Z9C] (complex with ''[[ohrA]]'' promoter) | ||
| Line 113: | Line 115: | ||
* '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=ohrR_1381434_1381877_-1 ohrR] {{PubMed|22383849}} | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=ohrR_1381434_1381877_-1 ohrR] {{PubMed|22383849}} | ||
| − | * '''Sigma factor:''' [[SigA]] | + | * '''[[Sigma factor]]:''' [[SigA]] |
* '''Regulation:''' | * '''Regulation:''' | ||
| Line 120: | Line 122: | ||
* '''Additional information:''' | * '''Additional information:''' | ||
| + | ** number of protein molecules per cell (minimal medium with glucose and ammonium): 133 {{PubMed|24696501}} | ||
=Biological materials = | =Biological materials = | ||
| Line 143: | Line 146: | ||
=References= | =References= | ||
| − | |||
==Reviews== | ==Reviews== | ||
| − | <pubmed>22797754 | + | <pubmed>22797754 20626317 19575568 20094649 25852656</pubmed> |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
==Original Publications== | ==Original Publications== | ||
| − | <pubmed>21749987,16209951,18487332,17660290,17502599,12486061,11418552,18586944,11983871,18363800,19129220 9696771 21749987 22797754 </pubmed> | + | <pubmed>21749987,16209951,18487332,17660290,17502599,12486061,11418552,18586944,11983871,18363800,19129220 9696771 21749987 22797754 24313874 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Latest revision as of 11:02, 10 April 2015
- Description: transcription repressor of the ohrA gene
| Gene name | ohrR |
| Synonyms | ykmA |
| Essential | no |
| Product | transcription repressor (MarR family) |
| Function | regulation of ohrA expression in response to organic peroxides |
| Gene expression levels in SubtiExpress: ohrR | |
| Interactions involving this protein in SubtInteract: OhrR | |
| MW, pI | 16 kDa, 6.364 |
| Gene length, protein length | 441 bp, 147 aa |
| Immediate neighbours | ohrA, ohrB |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 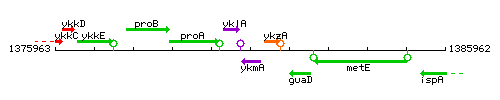
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed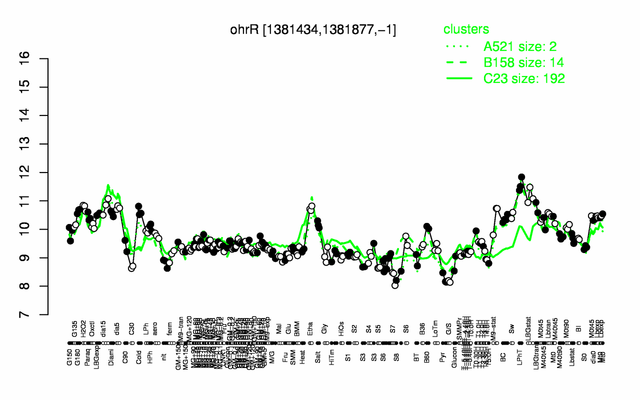
| |
Contents
Categories containing this gene/protein
transcription factors and their control, resistance against oxidative and electrophile stress
This gene is a member of the following regulons
The OhrR regulon:
The gene
Basic information
- Locus tag: BSU13150
Phenotypes of a mutant
Database entries
- BsubCyc: BSU13150
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: MarR family
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- Effectors of protein activity:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- BsubCyc: BSU13150
- UniProt: O34777
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: ohrR PubMed
- Regulation:
- Regulatory mechanism:
- Additional information:
- number of protein molecules per cell (minimal medium with glucose and ammonium): 133 PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
John Helmann, Cornell University, USA Homepage
Richard Brennan, Houston, Texas, USA Homepage
Your additional remarks
References
Reviews
Original Publications