Difference between revisions of "GlnA"
| Line 82: | Line 82: | ||
* '''Swiss prot entry:''' [http://www.uniprot.org/uniprot/P12425 P12425] | * '''Swiss prot entry:''' [http://www.uniprot.org/uniprot/P12425 P12425] | ||
| − | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu+BSU17460] | + | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu+BSU17460 BSU17460] |
* '''E.C. number:''' [http://www.expasy.org/enzyme/6.3.1.2 6.3.1.2] | * '''E.C. number:''' [http://www.expasy.org/enzyme/6.3.1.2 6.3.1.2] | ||
Revision as of 22:59, 13 May 2009
- Description: glutamine synthetase
| Gene name | glnA |
| Synonyms | |
| Essential | no |
| Product | trigger enzyme: glutamine synthetase |
| Function | glutamine biosynthesis, control of TnrA and GlnR activity |
| MW, pI | 50 kDa, 4.874 |
| Gene length, protein length | 1332 bp, 444 aa |
| Immediate neighbours | glnR, ynxB |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 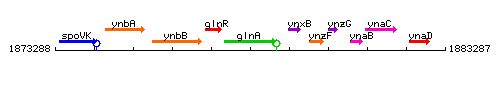
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Coordinates:
Phenotypes of a mutant
auxotrophic for glutamine
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information: K(M) for: Glu: 27 mM, ATP: 2.4 mM, ammonium: 0.18 mM; v(max): 3.7 µmol/min/mg
- Domains: glutamate binding flap (aa 300 ... 306: protects unstable intermediates from abberant hydrolysis)
- Modification: phosphorylated on ser/ thr/ tyr PubMed
- Cofactor(s): Mg(2+)
- Effectors of protein activity: feedback inhibition by glutamine, glutamine binds thhe entrance site for glutamate
- Interactions: TnrA-GlnA, GlnR-GlnA, (only the feedback-inhibited enzyme interacts with TnrA and GlnR)
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- Structure:
- Swiss prot entry: P12425
- KEGG entry: BSU17460
- E.C. number: 6.3.1.2
Additional information
GlnA is a homooligomer of 12 subunits
Expression and regulation
- Regulation: expressed in the absence of glutamine PubMed
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody: available in Karl Forchhammer lab
Labs working on this gene/protein
Susan Fisher, Boston, USA homepage
Your additional remarks
References
- Lévine et al. (2006) Analysis of the dynamic Bacillus subtilis Ser/Thr/Tyr phosphoproteome implicated in a wide variety of cellular processes. Proteomics 6: 2157-2173 PubMed
- Brown, S. W., and A. L. Sonenshein. 1996. Autogenous regulation of the Bacillus subtilis glnRA operon. J. Bacteriol. 178: 2450-2454. PubMed
- Wray LV Jr, Zalieckas JM, Fisher SH. (2001) Bacillus subtilis glutamine synthetase controls gene expression through protein-protein interaction with transcription factor TnrA. Cell 107:427-435. PubMed
- Fisher, S. H., and Wray, L. V., Jr. (2006) Feedback-resistant mutations in Bacillus subtilis glutamine synthetase are clustered in the active site. J Bacteriol 188: 5966-5974. PubMed
- Fisher SH, Sonenshein AL (1984) Bacillus subtilis glutamine synthetase mutants pleiotropically altered in catabolite repression. J Bacteriol 157:612-621. PubMed
- Fisher, S. H., Brandenburg, J. L. & Wray, L. V. (2002). Mutations in Bacillus subtilis glutamine synthetase that block its interaction with transcription factor TnrA. Mol Microbiol 45, 627-635. PubMed
- Schreier, H. J., Brown, S. W., Hirschi, K. D., Nomellini, J. F. & Sonenshein, A. L. (1989). Regulation of Bacillus subtilis glutamine synthetase gene expression by the product of the glnR gene. J Mol Biol 210, 51-63. PubMed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed