Difference between revisions of "CheY"
(→Expression and regulation) |
|||
| Line 57: | Line 57: | ||
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
| + | * not essential for pellicle biofilm formation, but mutant is outcompeted by the wild-type strain when competed during pellicle formation {{PubMed|26122431}} | ||
=== Database entries === | === Database entries === | ||
| Line 135: | Line 136: | ||
* '''Mutant:''' | * '''Mutant:''' | ||
** 1A798 ( ''cheY''::''cat''), {{PubMed|1905718}}, available at [http://pasture.asc.ohio-state.edu/BGSC/getdetail.cfm?bgscid=1A798&Search=1A798 BGSC] | ** 1A798 ( ''cheY''::''cat''), {{PubMed|1905718}}, available at [http://pasture.asc.ohio-state.edu/BGSC/getdetail.cfm?bgscid=1A798&Search=1A798 BGSC] | ||
| + | ** DS6870 (marker-less in NCIB3610) {{PubMed|25313396}} | ||
| + | ** TB191 ''amyE''::Phy-''sfgfp'' (marker-less in NCIB3610 with constitutive expressed ''sfgfp'') {{PubMed|26122431}} | ||
| + | ** TB207 ''amyE''::Phy-''mKATE2'' (marker-less in NCIB3610 with constitutive expressed ''mKATE2'') {{PubMed|26122431}} | ||
* '''Expression vector:''' | * '''Expression vector:''' | ||
| Line 152: | Line 156: | ||
=References= | =References= | ||
==Reviews== | ==Reviews== | ||
| − | <pubmed>19494578,,10094672 20122866 16224120 </pubmed> | + | <pubmed>25313396, 26122431, 19494578,,10094672 20122866 16224120 </pubmed> |
==Original Publications== | ==Original Publications== | ||
Revision as of 13:01, 2 July 2015
- Description: two-component response regulator, modulation of flagellar switch bias
| Gene name | cheY |
| Synonyms | cheB |
| Essential | no |
| Product | two-component response regulator |
| Function | modulation of flagellar switch bias |
| Gene expression levels in SubtiExpress: cheY | |
| Interactions involving this protein in SubtInteract: CheY | |
| MW, pI | 13 kDa, 4.746 |
| Gene length, protein length | 360 bp, 120 aa |
| Immediate neighbours | fliY, fliZ |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 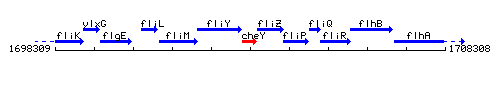
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed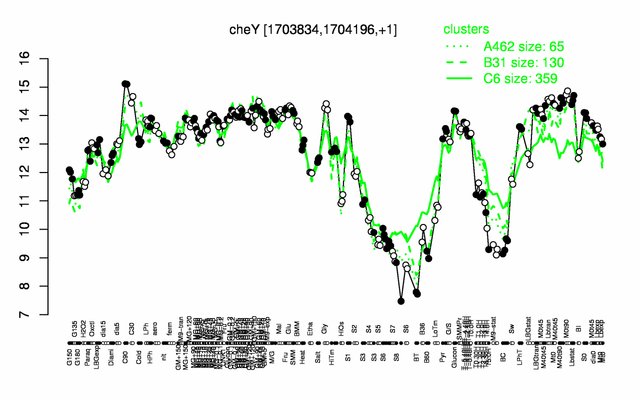
| |
Contents
Categories containing this gene/protein
transcription factors and their control, motility and chemotaxis, phosphoproteins
This gene is a member of the following regulons
CodY regulon, DegU regulon, SigD regulon, Spo0A regulon
The gene
Basic information
- Locus tag: BSU16330
Phenotypes of a mutant
- not essential for pellicle biofilm formation, but mutant is outcompeted by the wild-type strain when competed during pellicle formation PubMed
Database entries
- BsubCyc: BSU16330
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s): YneI
Extended information on the protein
- Kinetic information:
- Effectors of protein activity:
- Localization:
- cytoplasm (according to Swiss-Prot)
Database entries
- BsubCyc: BSU16330
- UniProt: P24072
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: part of the fla-che operon
- Regulation: see fla-che operon
- Regulatory mechanism: see fla-che operon
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Theresa Hölscher, Benjamin Bartels, Yu-Cheng Lin, Ramses Gallegos-Monterrosa, Alexa Price-Whelan, Roberto Kolter, Lars E P Dietrich, Ákos T Kovács
Motility, Chemotaxis and Aerotaxis Contribute to Competitiveness during Bacterial Pellicle Biofilm Development.
J Mol Biol: 2015, 427(23);3695-3708
[PubMed:26122431]
[WorldCat.org]
[DOI]
(I p)
Rebecca A Calvo, Daniel B Kearns
FlgM is secreted by the flagellar export apparatus in Bacillus subtilis.
J Bacteriol: 2015, 197(1);81-91
[PubMed:25313396]
[WorldCat.org]
[DOI]
(I p)
Gerald L Hazelbauer, Wing-Cheung Lai
Bacterial chemoreceptors: providing enhanced features to two-component signaling.
Curr Opin Microbiol: 2010, 13(2);124-32
[PubMed:20122866]
[WorldCat.org]
[DOI]
(I p)
Christopher V Rao, George W Ordal
The molecular basis of excitation and adaptation during chemotactic sensory transduction in bacteria.
Contrib Microbiol: 2009, 16;33-64
[PubMed:19494578]
[WorldCat.org]
[DOI]
(P p)
Christopher V Rao, John R Kirby, Adam P Arkin
Phosphatase localization in bacterial chemotaxis: divergent mechanisms, convergent principles.
Phys Biol: 2005, 2(3);148-58
[PubMed:16224120]
[WorldCat.org]
[DOI]
(I e)
C Fabret, V A Feher, J A Hoch
Two-component signal transduction in Bacillus subtilis: how one organism sees its world.
J Bacteriol: 1999, 181(7);1975-83
[PubMed:10094672]
[WorldCat.org]
[DOI]
(P p)
Original Publications