Difference between revisions of "LonA"
| Line 40: | Line 40: | ||
= [[Categories]] containing this gene/protein = | = [[Categories]] containing this gene/protein = | ||
| − | {{SubtiWiki category|[[proteolysis]]}} | + | {{SubtiWiki category|[[proteolysis]]}}, |
| + | {{SubtiWiki category|[[motility and chemotaxis]]}} | ||
= This gene is a member of the following [[regulons]] = | = This gene is a member of the following [[regulons]] = | ||
| Line 92: | Line 93: | ||
* '''[[SubtInteract|Interactions]]:''' | * '''[[SubtInteract|Interactions]]:''' | ||
** homohexameric protein {{PubMed|20600124}} | ** homohexameric protein {{PubMed|20600124}} | ||
| + | ** [[LonA]]-[[SmiA]] {{PubMed|25538299}} | ||
| + | ** [[LonA]]-[[SwrA]] (degradation) {{PubMed|25538299}} | ||
| − | * '''[[Localization]]:''' coincident with the nucleoid during normal growth and localized to the forespore during development {{PubMed|18689473}} | + | * '''[[Localization]]:''' |
| + | ** coincident with the nucleoid during normal growth and localized to the forespore during development {{PubMed|18689473}} | ||
=== Database entries === | === Database entries === | ||
| Line 148: | Line 152: | ||
==Original publications== | ==Original publications== | ||
| − | <pubmed>7961403,7961402, 19542270, 19643080 9852015 18689473 20600124 </pubmed> | + | <pubmed>7961403,7961402, 19542270, 19643080 9852015 18689473 20600124 25538299</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 11:18, 2 January 2015
- Description: class III heat-shock ATP-dependent protease
| Gene name | lonA |
| Synonyms | lon |
| Essential | no |
| Product | class III heat-shock ATP-dependent protease |
| Function | protein quality control |
| Gene expression levels in SubtiExpress: lonA | |
| Regulation of this protein in SubtiPathways: lonA | |
| MW, pI | 86 kDa, 5.774 |
| Gene length, protein length | 2322 bp, 774 aa |
| Immediate neighbours | ysxC, lonB |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 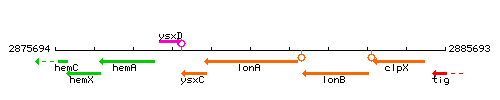
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed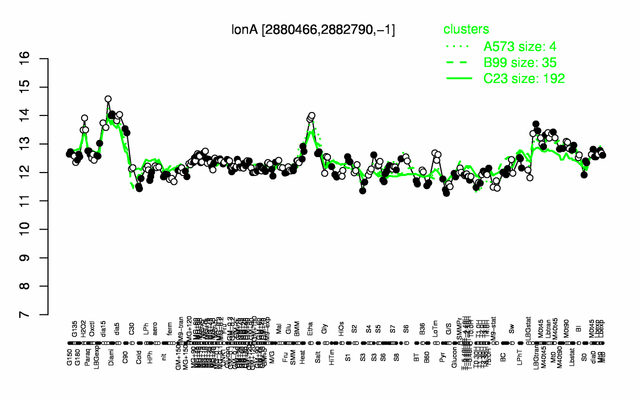
| |
Contents
Categories containing this gene/protein
proteolysis, motility and chemotaxis
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU28200
Phenotypes of a mutant
Database entries
- BsubCyc: BSU28200
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
A mutation in lonA suppresses the motility defect of a lytC mutant PubMed
Evidence for a specific DNA binding activitity of the α-domain was found PubMed
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Hydrolysis of proteins in presence of ATP (according to Swiss-Prot)
- Protein family: peptidase S16 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- coincident with the nucleoid during normal growth and localized to the forespore during development PubMed
Database entries
- BsubCyc: BSU28200
- UniProt: P37945
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications