Difference between revisions of "IleS"
| Line 126: | Line 126: | ||
* '''Additional information:''' subject to Clp-dependent proteolysis upon glucose starvation [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+17981983 PubMed] | * '''Additional information:''' subject to Clp-dependent proteolysis upon glucose starvation [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+17981983 PubMed] | ||
| + | ** number of protein molecules per cell (minimal medium with glucose and ammonium): 2829 {{PubMed|24696501}} | ||
| + | ** number of protein molecules per cell (complex medium with amino acids, without glucose): 9345 {{PubMed|24696501}} | ||
=Biological materials = | =Biological materials = | ||
Revision as of 09:45, 17 April 2014
- Description: isoleucyl-tRNA synthetase
| Gene name | ileS |
| Synonyms | |
| Essential | yes PubMed |
| Product | isoleucyl-tRNA synthetase |
| Function | translation |
| Gene expression levels in SubtiExpress: ileS | |
| Metabolic function and regulation of this protein in SubtiPathways: ileS | |
| MW, pI | 104 kDa, 5.191 |
| Gene length, protein length | 2763 bp, 921 aa |
| Immediate neighbours | divIVA, ylyA |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed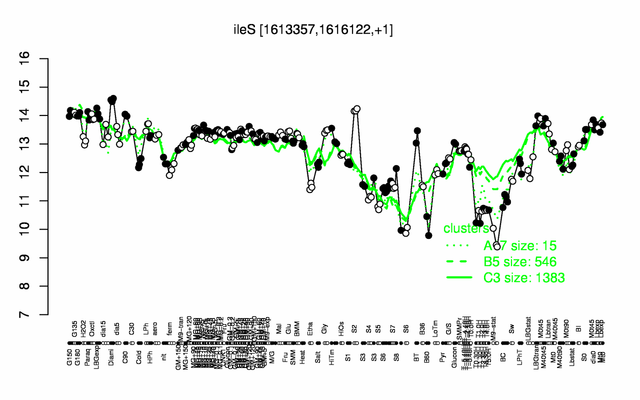
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU15430
Phenotypes of a mutant
essential PubMed
Database entries
- BsubCyc: BSU15430
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP + L-isoleucine + tRNA(Ile) = AMP + diphosphate + L-isoleucyl-tRNA(Ile) (according to Swiss-Prot)
- Protein family: IleS type 1 subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- cytoplasm (according to Swiss-Prot)
Database entries
- BsubCyc: BSU15430
- UniProt: Q45477
- KEGG entry: [2]
- E.C. number: 6.1.1.5
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Operon:
- Regulatory mechanism:
- T-box: RNA switch, transcriptional antitermination PubMed
- Additional information: subject to Clp-dependent proteolysis upon glucose starvation PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Ana Gutiérrez-Preciado, Tina M Henkin, Frank J Grundy, Charles Yanofsky, Enrique Merino
Biochemical features and functional implications of the RNA-based T-box regulatory mechanism.
Microbiol Mol Biol Rev: 2009, 73(1);36-61
[PubMed:19258532]
[WorldCat.org]
[DOI]
(I p)
Scott P Salowe, Judyann Wiltsie, Julio C Hawkins, Lisa M Sonatore
The catalytic flexibility of tRNAIle-lysidine synthetase can generate alternative tRNA substrates for isoleucyl-tRNA synthetase.
J Biol Chem: 2009, 284(15);9656-62
[PubMed:19233850]
[WorldCat.org]
[DOI]
(P p)
Ulf Gerth, Holger Kock, Ilja Kusters, Stephan Michalik, Robert L Switzer, Michael Hecker
Clp-dependent proteolysis down-regulates central metabolic pathways in glucose-starved Bacillus subtilis.
J Bacteriol: 2008, 190(1);321-31
[PubMed:17981983]
[WorldCat.org]
[DOI]
(I p)
L F Silvian, J Wang, T A Steitz
Insights into editing from an ile-tRNA synthetase structure with tRNAile and mupirocin.
Science: 1999, 285(5430);1074-7
[PubMed:10446055]
[WorldCat.org]
(P p)