Difference between revisions of "CheD"
(→Categories containing this gene/protein) |
|||
| Line 134: | Line 134: | ||
* '''Mutant:''' | * '''Mutant:''' | ||
| + | ** 1A861 ( ''cheD''::''cat''), {{PubMed|7893679}}, available at [http://pasture.asc.ohio-state.edu/BGSC/getdetail.cfm?bgscid=1A861&Search=1A861 BGSC] | ||
* '''Expression vector:''' | * '''Expression vector:''' | ||
Revision as of 09:00, 19 September 2012
- Description: protein deaminase, required for methylation of methyl-accepting chemotaxis proteins by CheR, enhances phosphatase activity of CheC, required for full McpC activity
| Gene name | cheD |
| Synonyms | ylxK |
| Essential | no |
| Product | protein deaminase, CheC activity modulator |
| Function | motility and chemotaxis |
| Gene expression levels in SubtiExpress: cheD | |
| Interactions involving this protein in SubtInteract: CheD | |
| MW, pI | 17 kDa, 9.347 |
| Gene length, protein length | 498 bp, 166 aa |
| Immediate neighbours | cheC, sigD |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 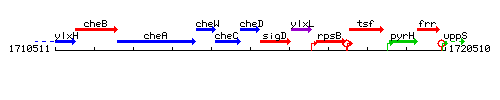
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed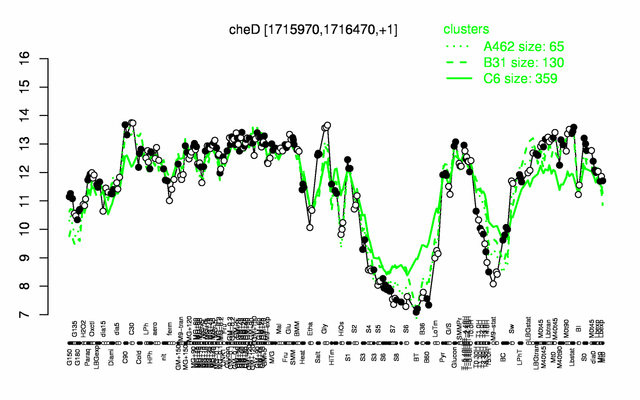
| |
Contents
Categories containing this gene/protein
protein modification, motility and chemotaxis
This gene is a member of the following regulons
CodY regulon, SigD regulon, Spo0A regulon
The gene
Basic information
- Locus tag: BSU16460
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: cheD family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: P40404
- KEGG entry: [3]
- E.C. number: 3.5.1.44
Additional information
Expression and regulation
- Operon:
- Regulatory mechanism:
- Additional information:
- in minimal medium, CheD is present with 1,200 +/- 250 molecules per cell PubMed
Biological materials
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Travis J Muff, George W Ordal
The diverse CheC-type phosphatases: chemotaxis and beyond.
Mol Microbiol: 2008, 70(5);1054-61
[PubMed:18990184]
[WorldCat.org]
[DOI]
(I p)
Christopher V Rao, George D Glekas, George W Ordal
The three adaptation systems of Bacillus subtilis chemotaxis.
Trends Microbiol: 2008, 16(10);480-7
[PubMed:18774298]
[WorldCat.org]
[DOI]
(P p)
Original publications