Difference between revisions of "CtsR"
(→Biological materials) |
(→Original publications) |
||
| Line 147: | Line 147: | ||
==Original publications== | ==Original publications== | ||
'''Additional publications:''' {{PubMed|17380125,16163393,19498169}} | '''Additional publications:''' {{PubMed|17380125,16163393,19498169}} | ||
| − | <pubmed>8793870, 9987115,12923084,11717291,16788169,11069659, 9852015,12884008,8195092,9987115,11544224 9852015, 11179229, 8793870 , 12884008 21208299 20852588 </pubmed> | + | <pubmed>8793870, 9987115,12923084,11717291,16788169,11069659, 9852015,12884008,8195092,9987115,11544224 9852015, 11179229, 8793870 , 12884008 21208299 20852588 22247503</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 09:08, 17 January 2012
| Gene name | ctsR |
| Synonyms | yacG |
| Essential | no |
| Product | transcription repressor |
| Function | regulation of protein degradation |
| Interactions involving this protein in SubtInteract: CtsR | |
| Regulatory function and regulation of this protein in SubtiPathways: Stress, Phosphorelay | |
| MW, pI | 17 kDa, 9.261 |
| Gene length, protein length | 462 bp, 154 aa |
| Immediate neighbours | rrnW-5S, mcsA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 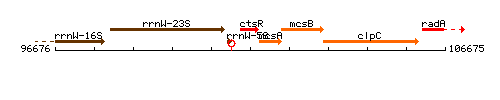
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
proteolysis, transcription factors and their control, general stress proteins (controlled by SigB), heat shock proteins, phosphoproteins
This gene is a member of the following regulons
The CtsR regulon
The gene
Basic information
- Locus tag: BSU00830
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: ctsR family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- UniProt: P37568
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Additional information: the mRNA is very stable (half-life > 15 min) PubMed
Biological materials
- Mutant: ctsR::aphA3 availbale from the Gerth lab
ctsRG65P::spec available from the Gerth lab
- Expression vector: for expression, purification in E. coli with N-terminal His-tag, pRSETA available in Gerth lab
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody: available in Gerth lab
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications
Additional publications: PubMed