Difference between revisions of "Pyk"
(→Extended information on the protein) |
|||
| Line 58: | Line 58: | ||
=== Additional information=== | === Additional information=== | ||
| − | + | * A mutation was found in this gene after evolution under relaxed selection for sporulation {{PubMed|21821766}} | |
| − | |||
=The protein= | =The protein= | ||
| Line 146: | Line 145: | ||
=References= | =References= | ||
| − | <pubmed>17726680 16493705 11489127 17505547 10966427 17218307 3711058 4623707 </pubmed> | + | <pubmed>17726680 16493705 11489127 17505547 10966427 17218307 3711058 4623707 21821766</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 11:46, 15 August 2011
- Description: pyruvate kinase, glycolytic enzyme
| Gene name | pyk |
| Synonyms | pykA |
| Essential | no |
| Product | pyruvate kinase |
| Function | catabolic enzyme in glycolysis |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 61,9 kDa, 4.88 |
| Gene length, protein length | 1755 bp, 585 amino acids |
| Immediate neighbours | ytzA, pfkA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 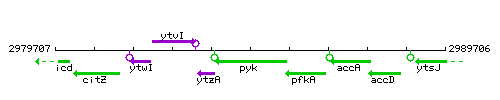
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
ATP synthesis, carbon core metabolism, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU29180
Phenotypes of a mutant
Unable to grow with non-PTS carbohydrates (such as glucitol or glycerol) as single carbon source.
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
- A mutation was found in this gene after evolution under relaxed selection for sporulation PubMed
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ADP + phosphoenolpyruvate --> ATP + pyruvate
- The reaction is irreversible under physiological conditions
- Protein family: PEP-utilizing enzyme family (according to Swiss-Prot) pyruvate kinase family, (C-terminal section: PEP-utilizing enzyme family)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information: Allosteric Regulation PubMed
- Domains:
- Modification: phosphorylation on Ser36 PubMed, PubMed, phosphorylation on Ser536 or Ser546 PubMed, please note that the Ser is not on position 536 but rather at 538
- Cofactor(s): Mg2+, K+
- Effectors of protein activity:
- Localization: cytoplasm PubMed
Database entries
- Structure: 2E28 (Geobacillus stearothermophilus)
- UniProt: P80885
- KEGG entry: [3]
- E.C. number: 2.7.1.40
Additional information
The enzyme is a tetramer with four active sites PubMed
Expression and regulation
- Sigma factor:
- Regulation:
- twofold induced by glucose PubMed
- Regulatory mechanism:
- Additional information:
Biological materials
- Expression vector:
- expression in E. coli, N-terminal His-tag: pGP1100 (in pWH844), available in Stülke lab
- expression in B. subtilis, native protein: pGP1411 (in pBQ200), available in Stülke lab
- expression in B. subtilis, N-terminal Strep-tag: pGP1409 (in pGP380), available in Stülke lab
- expression in B. subtilis, C-terminal Strep-tag: pGP1410 (in pGP382), available in Stülke lab
- lacZ fusion: see pfkA
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody:
Labs working on this gene/protein
Jörg Stülke, University of Göttingen, Germany Homepage
Your additional remarks
References