Difference between revisions of "OhrR"
| Line 74: | Line 74: | ||
* '''Domains:''' | * '''Domains:''' | ||
| − | * '''Modification:''' senses organic peroxides, released from DNA upon oxidation of an active site cysteine | + | * '''Modification:''' |
| + | ** senses organic peroxides, released from DNA upon oxidation of an active site cysteine | ||
| + | ** S-bacillithiolation upon hypochlorite stress on Cys-15, this results in release from the'' [[ohrA]]'' promoter {{PubMed|21749987}} | ||
* '''Cofactor(s):''' | * '''Cofactor(s):''' | ||
| Line 80: | Line 82: | ||
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| − | * '''Interactions:''' forms dimers | + | * '''[[SubtInteract|Interactions]]:''' |
| + | ** forms dimers | ||
* '''Localization:''' cytoplasm (according to Swiss-Prot) | * '''Localization:''' cytoplasm (according to Swiss-Prot) | ||
| Line 134: | Line 137: | ||
==Original Publications== | ==Original Publications== | ||
| − | <pubmed>16209951,18487332,17660290,17502599,12486061,11418552,18586944,11983871,18363800,19129220 9696771 </pubmed> | + | <pubmed>16209951,18487332,17660290,17502599,12486061,11418552,18586944,11983871,18363800,19129220 9696771 21749987 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 08:24, 14 July 2011
- Description: transcription repressor of the ohrA gene
| Gene name | ohrR |
| Synonyms | ykmA |
| Essential | no |
| Product | transcription repressor (MarR family) |
| Function | regulation of ohrA expression in response to organic peroxides |
| MW, pI | 16 kDa, 6.364 |
| Gene length, protein length | 441 bp, 147 aa |
| Immediate neighbours | ohrA, ohrB |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 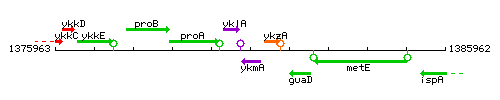
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
transcription factors and their control, resistance against oxidative and electrophile stress
This gene is a member of the following regulons
The OhrR regulon:
The gene
Basic information
- Locus tag: BSU13150
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- forms dimers
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- UniProt: O34777
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: ohrR PubMed
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
John Helmann, Cornell University, USA Homepage
Richard Brennan, Houston, Texas, USA Homepage
Your additional remarks
References
Reviews
Original Publications