Difference between revisions of "Mfd"
(→Original Articles) |
|||
| Line 13: | Line 13: | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || promotes strand-specific DNA repair by displacing | |style="background:#ABCDEF;" align="center"|'''Function''' || promotes strand-specific DNA repair by displacing | ||
| − | RNA polymerase stalled at a nucleotide lesion and directing | + | [[RNA polymerase]] stalled at a nucleotide lesion and directing |
the (A)BC excinuclease to the RNA damage site | the (A)BC excinuclease to the RNA damage site | ||
| Line 41: | Line 41: | ||
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
| − | |||
In an ''mfd'' knock-out, the cell's ability to accumulate adaptive mutations in stationary phase is depressed. [http://www.pubmed.com/16950921 PubMed] | In an ''mfd'' knock-out, the cell's ability to accumulate adaptive mutations in stationary phase is depressed. [http://www.pubmed.com/16950921 PubMed] | ||
Revision as of 17:39, 21 August 2010
- Description: transcription-repair coupling factor
| Gene name | mfd |
| Synonyms | |
| Essential | no |
| Product | transcription-repair coupling factor |
| Function | promotes strand-specific DNA repair by displacing
RNA polymerase stalled at a nucleotide lesion and directing the (A)BC excinuclease to the RNA damage site |
| MW, pI | 133 kDa, 5.367 |
| Gene length, protein length | 3531 bp, 1177 aa |
| Immediate neighbours | yabK, spoVT |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 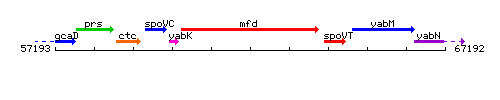
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU00550
Phenotypes of a mutant
In an mfd knock-out, the cell's ability to accumulate adaptive mutations in stationary phase is depressed. PubMed
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s): RecG
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
Database entries
- UniProt: P37474
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Operon:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant: GP1167 (ermC), available in Stülke lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original Articles