Difference between revisions of "YrrT"
(→Categories containing this gene/protein) |
|||
| Line 44: | Line 44: | ||
= [[Categories]] containing this gene/protein = | = [[Categories]] containing this gene/protein = | ||
| − | {{SubtiWiki category|[[poorly characterized/ putative enzymes]]}} | + | {{SubtiWiki category|[[poorly characterized/ putative enzymes]]}}, |
| + | {{SubtiWiki category|[[sporulation proteins]]}} | ||
= This gene is a member of the following [[regulons]] = | = This gene is a member of the following [[regulons]] = | ||
Revision as of 13:27, 8 May 2013
- Description: putative AdoMet-dependent methyltransferase
| Gene name | yrrT |
| Synonyms | |
| Essential | no |
| Product | putative AdoMet-dependent methyltransferase |
| Function | methionine salvage |
| Gene expression levels in SubtiExpress: yrrT | |
| Metabolic function and regulation of this protein in SubtiPathways: Cys, Met & Sulfate assimilation | |
| MW, pI | 24 kDa, 4.974 |
| Gene length, protein length | 639 bp, 213 aa |
| Immediate neighbours | mtnN, yrzA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 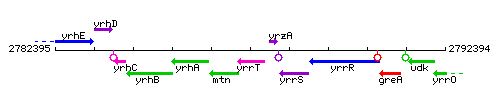
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed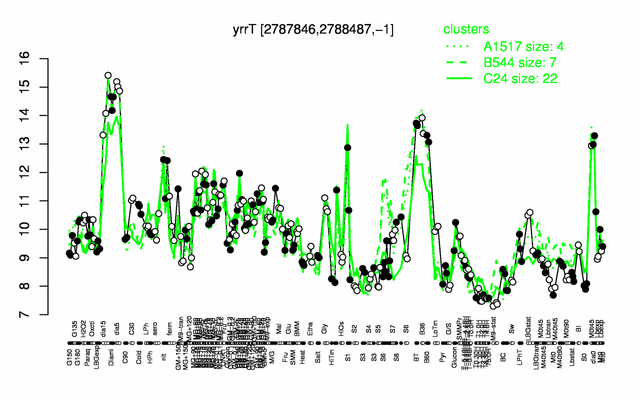
| |
Contents
Categories containing this gene/protein
poorly characterized/ putative enzymes, sporulation proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU27280
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure:
- UniProt: O32029
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Marie-Françoise Hullo, Sandrine Auger, Olga Soutourina, Octavian Barzu, Mireille Yvon, Antoine Danchin, Isabelle Martin-Verstraete
Conversion of methionine to cysteine in Bacillus subtilis and its regulation.
J Bacteriol: 2007, 189(1);187-97
[PubMed:17056751]
[WorldCat.org]
[DOI]
(P p)
Soon-Yong Choi, Dindo Reyes, Montira Leelakriangsak, Peter Zuber
The global regulator Spx functions in the control of organosulfur metabolism in Bacillus subtilis.
J Bacteriol: 2006, 188(16);5741-51
[PubMed:16885442]
[WorldCat.org]
[DOI]
(P p)
Sergine Even, Pierre Burguière, Sandrine Auger, Olga Soutourina, Antoine Danchin, Isabelle Martin-Verstraete
Global control of cysteine metabolism by CymR in Bacillus subtilis.
J Bacteriol: 2006, 188(6);2184-97
[PubMed:16513748]
[WorldCat.org]
[DOI]
(P p)
Shunji Nakano, Michiko M Nakano, Ying Zhang, Montira Leelakriangsak, Peter Zuber
A regulatory protein that interferes with activator-stimulated transcription in bacteria.
Proc Natl Acad Sci U S A: 2003, 100(7);4233-8
[PubMed:12642660]
[WorldCat.org]
[DOI]
(P p)