Difference between revisions of "SdhB"
Cadu Cunha (talk | contribs) (→Extended information on the protein) |
Cadu Cunha (talk | contribs) (→References) |
||
| Line 123: | Line 123: | ||
=References= | =References= | ||
| + | <pubmed> 3910107 6799760 </pubmed> | ||
# Hahne et al. (2008) From complementarity to comprehensiveness - targeting the membrane proteome of growing ''Bacillus subtilis'' by divergent approaches. Proteomics 8: 4123-4136 [http://www.ncbi.nlm.nih.gov/pubmed/18763711 PubMed] | # Hahne et al. (2008) From complementarity to comprehensiveness - targeting the membrane proteome of growing ''Bacillus subtilis'' by divergent approaches. Proteomics 8: 4123-4136 [http://www.ncbi.nlm.nih.gov/pubmed/18763711 PubMed] | ||
# Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | # Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | ||
Revision as of 10:43, 11 June 2009
- Description: succinate dehydrogenase
| Gene name | sdhB |
| Synonyms | |
| Essential | no |
| Product | succinate dehydrogenase (iron-sulfur protein) |
| Function | TCA cycle |
| MW, pI | 28 kDa, 7.989 |
| Gene length, protein length | 759 bp, 253 aa |
| Immediate neighbours | sdhA, ysmA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 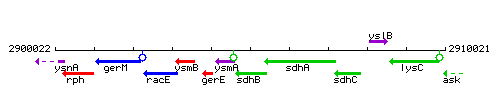
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU28430
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Succinate + acceptor = fumarate + reduced acceptor (according to Swiss-Prot)
- Protein family: succinate dehydrogenase/fumarate reductase iron-sulfur protein family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s): Fe
- Effectors of protein activity:
- Localization: attached to the membrane PubMed
Database entries
- Structure: 1NEK (E. coli)
- Swiss prot entry: P08066
- KEGG entry: [3]
- E.C. number:EC 1.3.99.1
Additional information
This enzyme is a trimer membrane-bound PubMed PubMed
- One subunit is bound to citochrome b558, and this subunit is the one bound to the cytosolic side of the membrane PubMed PubMed
- Another subunit is the flavoprotein one, required for FAD usage PubMed PubMed
- The other subunit has an iron-sulphur domain necessary for the catalytic activity PubMed PubMed
Expression and regulation
- Regulation: constitutive
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
S T Cole, C Condon, B D Lemire, J H Weiner
Molecular biology, biochemistry and bioenergetics of fumarate reductase, a complex membrane-bound iron-sulfur flavoenzyme of Escherichia coli.
Biochim Biophys Acta: 1985, 811(4);381-403
[PubMed:3910107]
[WorldCat.org]
[DOI]
(P p)
L Hederstedt, L Rutberg
Succinate dehydrogenase--a comparative review.
Microbiol Rev: 1981, 45(4);542-55
[PubMed:6799760]
[WorldCat.org]
[DOI]
(P p)