Difference between revisions of "RadC"
m (→Your additional remarks) |
|||
| Line 145: | Line 145: | ||
=Your additional remarks= | =Your additional remarks= | ||
| + | The ''radC'' locus is the chromosomal integration site for the φ105 prophage [http://www.ncbi.nlm.nih.gov/pubmed/24009114 Pubmed] | ||
=References= | =References= | ||
Revision as of 01:39, 8 June 2014
- Description: probable DNA repair protein
| Gene name | radC |
| Synonyms | ysxA |
| Essential | no |
| Product | probable DNA repair protein |
| Function | DNA repair |
| Gene expression levels in SubtiExpress: radC | |
| MW, pI | 25 kDa, 7.829 |
| Gene length, protein length | 693 bp, 231 aa |
| Immediate neighbours | mreB, maf |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 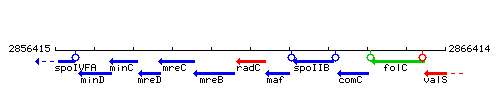
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed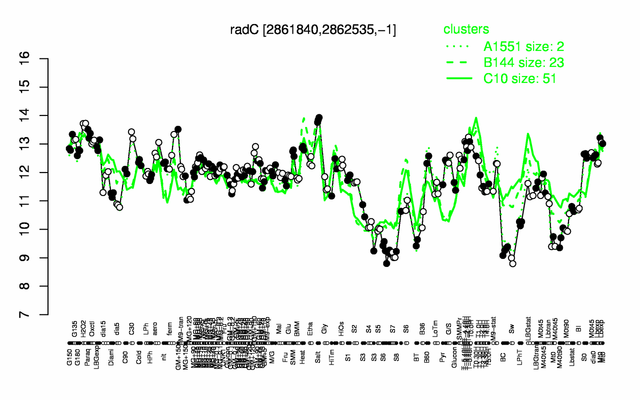
| |
Contents
Categories containing this gene/protein
DNA repair/ recombination, genetic competence, cell envelope stress proteins (controlled by SigM, V, W, X, Y), phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU28040
Phenotypes of a mutant
Database entries
- BsubCyc: BSU28040
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: radC family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- phosphorylated on Arg-59 PubMed
- Cofactor(s):
- Effectors of protein activity:
- Localization: Cytoplasm (Homogeneous) PubMed
Database entries
- BsubCyc: BSU28040
- Structure:
- UniProt: Q02170
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
The radC locus is the chromosomal integration site for the φ105 prophage Pubmed
References
Additional publications: PubMed
Alexander K W Elsholz, Kürsad Turgay, Stephan Michalik, Bernd Hessling, Katrin Gronau, Dan Oertel, Ulrike Mäder, Jörg Bernhardt, Dörte Becher, Michael Hecker, Ulf Gerth
Global impact of protein arginine phosphorylation on the physiology of Bacillus subtilis.
Proc Natl Acad Sci U S A: 2012, 109(19);7451-6
[PubMed:22517742]
[WorldCat.org]
[DOI]
(I p)
Warawan Eiamphungporn, John D Helmann
The Bacillus subtilis sigma(M) regulon and its contribution to cell envelope stress responses.
Mol Microbiol: 2008, 67(4);830-48
[PubMed:18179421]
[WorldCat.org]
[DOI]
(P p)
Adrian J Jervis, Penny D Thackray, Chris W Houston, Malcolm J Horsburgh, Anne Moir
SigM-responsive genes of Bacillus subtilis and their promoters.
J Bacteriol: 2007, 189(12);4534-8
[PubMed:17434969]
[WorldCat.org]
[DOI]
(P p)
Jean-Christophe Meile, Ling Juan Wu, S Dusko Ehrlich, Jeff Errington, Philippe Noirot
Systematic localisation of proteins fused to the green fluorescent protein in Bacillus subtilis: identification of new proteins at the DNA replication factory.
Proteomics: 2006, 6(7);2135-46
[PubMed:16479537]
[WorldCat.org]
[DOI]
(P p)
Penny D Thackray, Anne Moir
SigM, an extracytoplasmic function sigma factor of Bacillus subtilis, is activated in response to cell wall antibiotics, ethanol, heat, acid, and superoxide stress.
J Bacteriol: 2003, 185(12);3491-8
[PubMed:12775685]
[WorldCat.org]
[DOI]
(P p)
Mitsuo Ogura, Hirotake Yamaguchi, Kazuo Kobayashi, Naotake Ogasawara, Yasutaro Fujita, Teruo Tanaka
Whole-genome analysis of genes regulated by the Bacillus subtilis competence transcription factor ComK.
J Bacteriol: 2002, 184(9);2344-51
[PubMed:11948146]
[WorldCat.org]
[DOI]
(P p)
Randy M Berka, Jeanette Hahn, Mark Albano, Irena Draskovic, Marjan Persuh, Xianju Cui, Alan Sloma, William Widner, David Dubnau
Microarray analysis of the Bacillus subtilis K-state: genome-wide expression changes dependent on ComK.
Mol Microbiol: 2002, 43(5);1331-45
[PubMed:11918817]
[WorldCat.org]
[DOI]
(P p)