Difference between revisions of "PyrG"
| Line 82: | Line 82: | ||
* '''Structure:''' | * '''Structure:''' | ||
| − | * ''' | + | * '''UniProt:''' [http://www.uniprot.org/uniprot/P13242 P13242] |
* '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu:BSU37150] | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu:BSU37150] | ||
Revision as of 15:00, 20 July 2009
- Description: CTP synthase (NH3, glutamine)
| Gene name | pyrG |
| Synonyms | ctrA |
| Essential | yes PubMed |
| Product | CTP synthase (NH3, glutamine) |
| Function | pyrimidine biosynthesis |
| Metabolic function and regulation of this protein in SubtiPathways: Pyrimidines, Nucleotides (regulation) | |
| MW, pI | 59 kDa, 5.16 |
| Gene length, protein length | 1605 bp, 535 aa |
| Immediate neighbours | ywjG, rpoE |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 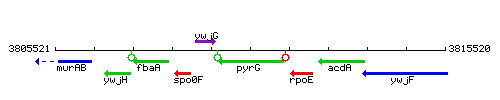
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU37150
Phenotypes of a mutant
essential PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP + UTP + NH3 = ADP + phosphate + CTP (according to Swiss-Prot)
- Protein family: CTP synthase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure:
- UniProt: P13242
- KEGG entry: [3]
- E.C. number: 6.3.4.2
Additional information
Expression and regulation
- Operon:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Irene E Jensen-MacAllister, Qi Meng, Robert L Switzer
Regulation of pyrG expression in Bacillus subtilis: CTP-regulated antitermination and reiterative transcription with pyrG templates in vitro.
Mol Microbiol: 2007, 63(5);1440-52
[PubMed:17302819]
[WorldCat.org]
[DOI]
(P p)
Alexander K W Elsholz, Casper Møller Jørgensen, Robert L Switzer
The number of G residues in the Bacillus subtilis pyrG initially transcribed region governs reiterative transcription-mediated regulation.
J Bacteriol: 2007, 189(5);2176-80
[PubMed:17158658]
[WorldCat.org]
[DOI]
(P p)
Qi Meng, Charles L Turnbough, Robert L Switzer
Attenuation control of pyrG expression in Bacillus subtilis is mediated by CTP-sensitive reiterative transcription.
Proc Natl Acad Sci U S A: 2004, 101(30);10943-8
[PubMed:15252202]
[WorldCat.org]
[DOI]
(P p)
Qi Meng, Robert L Switzer
cis-acting sequences of Bacillus subtilis pyrG mRNA essential for regulation by antitermination.
J Bacteriol: 2002, 184(23);6734-8
[PubMed:12426364]
[WorldCat.org]
[DOI]
(P p)
Q Meng, R L Switzer
Regulation of transcription of the Bacillus subtilis pyrG gene, encoding cytidine triphosphate synthetase.
J Bacteriol: 2001, 183(19);5513-22
[PubMed:11544212]
[WorldCat.org]
[DOI]
(P p)
R J Turner, Y Lu, R L Switzer
Regulation of the Bacillus subtilis pyrimidine biosynthetic (pyr) gene cluster by an autogenous transcriptional attenuation mechanism.
J Bacteriol: 1994, 176(12);3708-22
[PubMed:8206849]
[WorldCat.org]
[DOI]
(P p)
C L Quinn, B T Stephenson, R L Switzer
Functional organization and nucleotide sequence of the Bacillus subtilis pyrimidine biosynthetic operon.
J Biol Chem: 1991, 266(14);9113-27
[PubMed:1709162]
[WorldCat.org]
(P p)