Difference between revisions of "OdhB"
(→Extended information on the protein) |
(→References) |
||
| Line 143: | Line 143: | ||
=References= | =References= | ||
| − | + | ==Reviews== | |
| + | <pubmed> 10672230</pubmed> | ||
| + | ==Original publications== | ||
<pubmed>2500417,,1508153,12850135 18763711 12682299,11976317 20933603</pubmed> | <pubmed>2500417,,1508153,12850135 18763711 12682299,11976317 20933603</pubmed> | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 11:28, 26 December 2011
- Description: 2-oxoglutarate dehydrogenase complex (dihydrolipoamide transsuccinylase, E2 subunit)
| Gene name | odhB |
| Synonyms | citM |
| Essential | yes PubMed |
| Product | 2-oxoglutarate dehydrogenase complex
(dihydrolipoamide transsuccinylase, E2 subunit) |
| Function | TCA cycle |
| Interactions involving this protein in SubtInteract: OdhB | |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 45 kDa, 4.859 |
| Gene length, protein length | 1251 bp, 417 aa |
| Immediate neighbours | yocS, odhA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 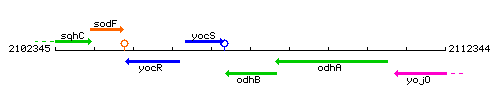
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
carbon core metabolism, essential genes, membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU19360
Phenotypes of a mutant
essential PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Succinyl-CoA + enzyme N(6)-(dihydrolipoyl)lysine = CoA + enzyme N(6)-(S-succinyldihydrolipoyl)lysine (according to Swiss-Prot)
- Protein family: sodium:bile acid symporter family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- membrane associated PubMed
Database entries
- UniProt: P16263
- KEGG entry: [3]
- E.C. number: 2.3.1.61
Additional information
- extensive information on the structure and enzymatic properties of 2-oxoglutarate dehydrogenase can be found at Proteopedia
Expression and regulation
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
K F Sheu, J P Blass
The alpha-ketoglutarate dehydrogenase complex.
Ann N Y Acad Sci: 1999, 893;61-78
[PubMed:10672230]
[WorldCat.org]
[DOI]
(P p)
Original publications
Frederik M Meyer, Jan Gerwig, Elke Hammer, Christina Herzberg, Fabian M Commichau, Uwe Völker, Jörg Stülke
Physical interactions between tricarboxylic acid cycle enzymes in Bacillus subtilis: evidence for a metabolon.
Metab Eng: 2011, 13(1);18-27
[PubMed:20933603]
[WorldCat.org]
[DOI]
(I p)
Hannes Hahne, Susanne Wolff, Michael Hecker, Dörte Becher
From complementarity to comprehensiveness--targeting the membrane proteome of growing Bacillus subtilis by divergent approaches.
Proteomics: 2008, 8(19);4123-36
[PubMed:18763711]
[WorldCat.org]
[DOI]
(I p)
Hans-Matti Blencke, Georg Homuth, Holger Ludwig, Ulrike Mäder, Michael Hecker, Jörg Stülke
Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways.
Metab Eng: 2003, 5(2);133-49
[PubMed:12850135]
[WorldCat.org]
[DOI]
(P p)
K Kobayashi, S D Ehrlich, A Albertini, G Amati, K K Andersen, M Arnaud, K Asai, S Ashikaga, S Aymerich, P Bessieres, F Boland, S C Brignell, S Bron, K Bunai, J Chapuis, L C Christiansen, A Danchin, M Débarbouille, E Dervyn, E Deuerling, K Devine, S K Devine, O Dreesen, J Errington, S Fillinger, S J Foster, Y Fujita, A Galizzi, R Gardan, C Eschevins, T Fukushima, K Haga, C R Harwood, M Hecker, D Hosoya, M F Hullo, H Kakeshita, D Karamata, Y Kasahara, F Kawamura, K Koga, P Koski, R Kuwana, D Imamura, M Ishimaru, S Ishikawa, I Ishio, D Le Coq, A Masson, C Mauël, R Meima, R P Mellado, A Moir, S Moriya, E Nagakawa, H Nanamiya, S Nakai, P Nygaard, M Ogura, T Ohanan, M O'Reilly, M O'Rourke, Z Pragai, H M Pooley, G Rapoport, J P Rawlins, L A Rivas, C Rivolta, A Sadaie, Y Sadaie, M Sarvas, T Sato, H H Saxild, E Scanlan, W Schumann, J F M L Seegers, J Sekiguchi, A Sekowska, S J Séror, M Simon, P Stragier, R Studer, H Takamatsu, T Tanaka, M Takeuchi, H B Thomaides, V Vagner, J M van Dijl, K Watabe, A Wipat, H Yamamoto, M Yamamoto, Y Yamamoto, K Yamane, K Yata, K Yoshida, H Yoshikawa, U Zuber, N Ogasawara
Essential Bacillus subtilis genes.
Proc Natl Acad Sci U S A: 2003, 100(8);4678-83
[PubMed:12682299]
[WorldCat.org]
[DOI]
(P p)
Ciarán Condon, Jordi Rourera, Dominique Brechemier-Baey, Harald Putzer
Ribonuclease M5 has few, if any, mRNA substrates in Bacillus subtilis.
J Bacteriol: 2002, 184(10);2845-9
[PubMed:11976317]
[WorldCat.org]
[DOI]
(P p)
O Resnekov, L Melin, P Carlsson, M Mannerlöv, A von Gabain, L Hederstedt
Organization and regulation of the Bacillus subtilis odhAB operon, which encodes two of the subenzymes of the 2-oxoglutarate dehydrogenase complex.
Mol Gen Genet: 1992, 234(2);285-96
[PubMed:1508153]
[WorldCat.org]
[DOI]
(P p)
P Carlsson, L Hederstedt
Genetic characterization of Bacillus subtilis odhA and odhB, encoding 2-oxoglutarate dehydrogenase and dihydrolipoamide transsuccinylase, respectively.
J Bacteriol: 1989, 171(7);3667-72
[PubMed:2500417]
[WorldCat.org]
[DOI]
(P p)