LeuD
- Description: 3-isopropylmalate dehydratase (small subunit)
| Gene name | leuD |
| Synonyms | |
| Essential | no |
| Product | 3-isopropylmalate dehydratase (small subunit) |
| Function | biosynthesis of leucine |
| Metabolic function and regulation of this protein in SubtiPathways: Ile, Leu, Val | |
| MW, pI | 22 kDa, 4.582 |
| Gene length, protein length | 597 bp, 199 aa |
| Immediate neighbours | ysoA, leuC |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 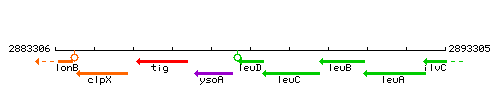
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU28250
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: (2R,3S)-3-isopropylmalate = (2S)-2-isopropylmaleate + H2O (according to Swiss-Prot)
- Protein family: LeuD type 1 subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure:
- Swiss prot entry: P94568
- KEGG entry: [3]
- E.C. number: 4.2.1.33
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Regulation: repressed by casamino acids PubMed , expressed in the absence of branched-chain amino acids (BCAA), expression is stimulated in the presence of glucose PubMed, repressed by CodY PubMed
- Regulatory mechanism: CodY: transcription repression PubMed, glucose regulation: CcpA PubMed, repression by BCAA: tRNA-controlled RNA switch (T-box) that mediates termination/antitermination
- Additional information: subject to Clp-dependent proteolysis upon glucose starvation PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Ana Gutiérrez-Preciado, Tina M Henkin, Frank J Grundy, Charles Yanofsky, Enrique Merino
Biochemical features and functional implications of the RNA-based T-box regulatory mechanism.
Microbiol Mol Biol Rev: 2009, 73(1);36-61
[PubMed:19258532]
[WorldCat.org]
[DOI]
(I p)
Ahmed Gaballa, Haike Antelmann, Claudio Aguilar, Sukhjit K Khakh, Kyung-Bok Song, Gregory T Smaldone, John D Helmann
The Bacillus subtilis iron-sparing response is mediated by a Fur-regulated small RNA and three small, basic proteins.
Proc Natl Acad Sci U S A: 2008, 105(33);11927-32
[PubMed:18697947]
[WorldCat.org]
[DOI]
(I p)
Shigeo Tojo, Takenori Satomura, Kanako Kumamoto, Kazutake Hirooka, Yasutaro Fujita
Molecular mechanisms underlying the positive stringent response of the Bacillus subtilis ilv-leu operon, involved in the biosynthesis of branched-chain amino acids.
J Bacteriol: 2008, 190(18);6134-47
[PubMed:18641142]
[WorldCat.org]
[DOI]
(I p)
Shigeo Tojo, Takenori Satomura, Kaori Morisaki, Ken-Ichi Yoshida, Kazutake Hirooka, Yasutaro Fujita
Negative transcriptional regulation of the ilv-leu operon for biosynthesis of branched-chain amino acids through the Bacillus subtilis global regulator TnrA.
J Bacteriol: 2004, 186(23);7971-9
[PubMed:15547269]
[WorldCat.org]
[DOI]
(P p)
Ulrike Mäder, Susanne Hennig, Michael Hecker, Georg Homuth
Transcriptional organization and posttranscriptional regulation of the Bacillus subtilis branched-chain amino acid biosynthesis genes.
J Bacteriol: 2004, 186(8);2240-52
[PubMed:15060025]
[WorldCat.org]
[DOI]
(P p)
Virginie Molle, Yoshiko Nakaura, Robert P Shivers, Hirotake Yamaguchi, Richard Losick, Yasutaro Fujita, Abraham L Sonenshein
Additional targets of the Bacillus subtilis global regulator CodY identified by chromatin immunoprecipitation and genome-wide transcript analysis.
J Bacteriol: 2003, 185(6);1911-22
[PubMed:12618455]
[WorldCat.org]
[DOI]
(P p)
Holger Ludwig, Christoph Meinken, Anastasija Matin, Jörg Stülke
Insufficient expression of the ilv-leu operon encoding enzymes of branched-chain amino acid biosynthesis limits growth of a Bacillus subtilis ccpA mutant.
J Bacteriol: 2002, 184(18);5174-8
[PubMed:12193635]
[WorldCat.org]
[DOI]
(P p)
Ulrike Mäder, Georg Homuth, Christian Scharf, Knut Büttner, Rüdiger Bode, Michael Hecker
Transcriptome and proteome analysis of Bacillus subtilis gene expression modulated by amino acid availability.
J Bacteriol: 2002, 184(15);4288-95
[PubMed:12107147]
[WorldCat.org]
[DOI]
(P p)
Christine Eymann, Georg Homuth, Christian Scharf, Michael Hecker
Bacillus subtilis functional genomics: global characterization of the stringent response by proteome and transcriptome analysis.
J Bacteriol: 2002, 184(9);2500-20
[PubMed:11948165]
[WorldCat.org]
[DOI]
(P p)
F J Grundy, T M Henkin
Conservation of a transcription antitermination mechanism in aminoacyl-tRNA synthetase and amino acid biosynthesis genes in gram-positive bacteria.
J Mol Biol: 1994, 235(2);798-804
[PubMed:8289305]
[WorldCat.org]
[DOI]
(P p)