Difference between revisions of "KtrB"
| Line 83: | Line 83: | ||
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| − | * '''Interactions:''' [[KtrA]]-[[KtrB]] [http://www.ncbi.nlm.nih.gov/sites/entrez/12562800 PubMed] | + | * '''Interactions:''' |
| + | ** KtrB forms dimers {{PubMed|17932047}} | ||
| + | ** [[KtrA]]-[[KtrB]] [http://www.ncbi.nlm.nih.gov/sites/entrez/12562800 PubMed] | ||
* '''Localization:''' cell membrane (according to Swiss-Prot), integral membrane protein [http://www.ncbi.nlm.nih.gov/sites/entrez/12562800 PubMed] | * '''Localization:''' cell membrane (according to Swiss-Prot), integral membrane protein [http://www.ncbi.nlm.nih.gov/sites/entrez/12562800 PubMed] | ||
| Line 134: | Line 136: | ||
=References= | =References= | ||
| − | <pubmed>12562800,20511502, </pubmed> | + | <pubmed>12562800,20511502, 17932047 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 11:10, 26 December 2010
| Gene name | ktrB |
| Synonyms | yubG |
| Essential | no |
| Product | high affinity potassium transporter KtrA-KtrB, integral membrane subunit (proton symport) |
| Function | potassium uptake |
| Metabolic function and regulation of this protein in SubtiPathways: Stress | |
| MW, pI | 48 kDa, 10.116 |
| Gene length, protein length | 1335 bp, 445 aa |
| Immediate neighbours | ktrA, yubF |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 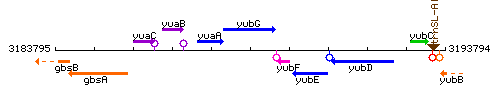
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
transporters/ other, metal ion homeostasis (K, Na, Ca, Mg), coping with hyper-osmotic stress, membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU31100
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s): KtrD
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: cell membrane (according to Swiss-Prot), integral membrane protein PubMed
Database entries
- Structure:
- UniProt: O32081
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- expression is controlled via termination antitermination by the ydaO riboswitch PubMed
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Erhard Bremer, University of Marburg, Germany homepage
Your additional remarks
References
Kirsten F Block, Ming C Hammond, Ronald R Breaker
Evidence for widespread gene control function by the ydaO riboswitch candidate.
J Bacteriol: 2010, 192(15);3983-9
[PubMed:20511502]
[WorldCat.org]
[DOI]
(I p)
Ronald A Albright, Kyu Joh, João H Morais-Cabral
Probing the structure of the dimeric KtrB membrane protein.
J Biol Chem: 2007, 282(48);35046-55
[PubMed:17932047]
[WorldCat.org]
[DOI]
(P p)
Gudrun Holtmann, Evert P Bakker, Nobuyuki Uozumi, Erhard Bremer
KtrAB and KtrCD: two K+ uptake systems in Bacillus subtilis and their role in adaptation to hypertonicity.
J Bacteriol: 2003, 185(4);1289-98
[PubMed:12562800]
[WorldCat.org]
[DOI]
(P p)