GudB
- Description: trigger enzyme: glutamate dehydrogenase (cryptic in 168 and derivatives)
| Gene name | gudB |
| Synonyms | ypcA |
| Essential | no |
| Product | trigger enzyme: glutamate dehydrogenase |
| Function | glutamate utilization, control of GltC activity |
| Gene expression levels in SubtiExpress: gudB | |
| Interactions involving this protein in SubtInteract: GudB | |
| Metabolic function and regulation of this protein in SubtiPathways: gudB | |
| MW, pI | 47 kDa, 5.582 |
| Gene length, protein length | 1278 bp, 426 aa |
| Immediate neighbours | ypdA, ypbH |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 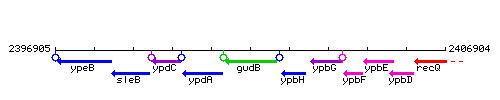
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed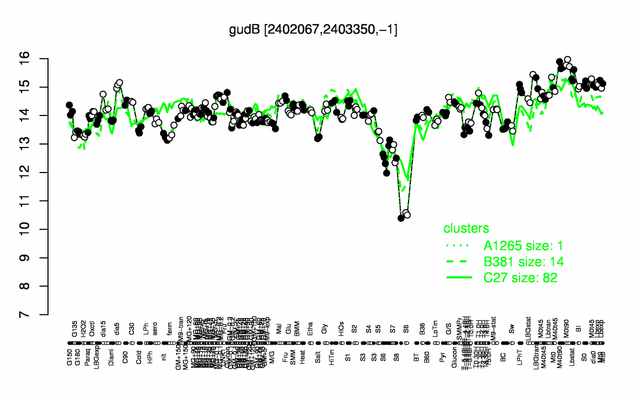
| |
Contents
Categories containing this gene/protein
utilization of amino acids, glutamate metabolism, transcription factors and their control, trigger enzyme, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU22960
Phenotypes of a mutant
- The gene is cryptic. If gudB is activated (gudB1 mutation), the bacteria are able to utilize glutamate as the only carbon source. PubMed
- A rocG gudB mutant is sensitive to ß-lactam antibiotics such as cefuroxime and to fosfomycin due to the downregulation of the SigW regulon PubMed
- transcription profile of a rocG gudB mutant strain: GEO PubMed
Database entries
- BsubCyc: BSU22960
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: L-glutamate + H2O + NAD+ = 2-oxoglutarate + NH3 + NADH + H+ (according to Swiss-Prot)
- Protein family: Glu/Leu/Phe/Val dehydrogenases family (according to Swiss-Prot)
- Paralogous protein(s): RocG
Extended information on the protein
- Kinetic information:
- Modification:
- phosphorylated on Arg-56, Arg-83, and Arg-421 and/or Arg-423 PubMed
- Cofactors: NAD+/NADH + H+
- Effectors of protein activity:
Database entries
- BsubCyc: BSU22960
- UniProt: P50735
- KEGG entry: [4]
- E.C. number: 1.4.1.2
Additional information
Expression and regulation
- Operon: gudB PubMed
- Regulation: constitutively expressed PubMed
- Regulatory mechanism:
- Additional information: GudB is subject to Clp-dependent proteolysis upon glucose starvation PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium): 239 PubMed
- number of protein molecules per cell (complex medium with amino acids, without glucose): 6565 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, exponential phase): 1157 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, early stationary phase after glucose exhaustion): 580 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, late stationary phase after glucose exhaustion): 888 PubMed
Biological materials
- Mutant: GP691 (ΔgudB::cat), GP1160 (ΔgudB::aphA3) both available in Jörg Stülke's lab
- Mutant: BP442 (ΔgudB::aphA3), lacking the complete promoter, available in Fabian Commichau's lab
- Expression vector:
- for purification of GudB from E. coli carrying an N-terminal Strep-tag: pGP863 (in pGP172) available in Jörg Stülke's lab
- for purification of GudB1 from E. coli carrying an N-terminal Strep-tag: pGP864 (in pGP172) available in Jörg Stülke's lab
- for ectopic expression of gudB with its native promoter: pGP900 (in pAC5), available in Jörg Stülke's lab
- wild type gudB, expression in B. subtilis, in pBQ200: pGP1712, available in Jörg Stülke's lab
- lacZ fusion: pGP651 (in pAC5), available in Jörg Stülke's lab
- FLAG-tag construct: GP1194 (gudB, spc, based on pGP1331), GP1195 (gudB1, spc, based on pGP1331), available in Jörg Stülke's lab
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH) PubMed, available in Jörg Stülke's lab
- Antibody: antibody against RocG recognizes GudB, available in Jörg Stülke's lab
Labs working on this gene/protein
- Linc Sonenshein, Tufts University, Boston, MA, USA Homepage
- Fabian Commichau University of Göttingen, Germany Homepage
Your additional remarks
The GudB protein is active in other legacy B. subtilis strains (e.g. strain 122). Thus, it can be speculated that the ancestral gudB gene was not cryptic, but became so as a product of the "domestication" of B. subtilis 168 in the lab. PubMed
References
Reviews
Original publications
Lorena Stannek, Martin J Thiele, Till Ischebeck, Katrin Gunka, Elke Hammer, Uwe Völker, Fabian M Commichau
Evidence for synergistic control of glutamate biosynthesis by glutamate dehydrogenases and glutamate in Bacillus subtilis.
Environ Microbiol: 2015, 17(9);3379-90
[PubMed:25711804]
[WorldCat.org]
[DOI]
(I p)
Lorena Stannek, Katrin Gunka, Rachel A Care, Ulf Gerth, Fabian M Commichau
Factors that mediate and prevent degradation of the inactive and unstable GudB protein in Bacillus subtilis.
Front Microbiol: 2014, 5;758
[PubMed:25610436]
[WorldCat.org]
[DOI]
(P e)
Lorena Stannek, Richard Egelkamp, Katrin Gunka, Fabian M Commichau
Monitoring intraspecies competition in a bacterial cell population by cocultivation of fluorescently labelled strains.
J Vis Exp: 2014, (83);e51196
[PubMed:24473333]
[WorldCat.org]
[DOI]
(I e)
Andreas Schmidt, Débora Broch Trentini, Silvia Spiess, Jakob Fuhrmann, Gustav Ammerer, Karl Mechtler, Tim Clausen
Quantitative phosphoproteomics reveals the role of protein arginine phosphorylation in the bacterial stress response.
Mol Cell Proteomics: 2014, 13(2);537-50
[PubMed:24263382]
[WorldCat.org]
[DOI]
(I p)
Katrin Gunka, Lorena Stannek, Rachel A Care, Fabian M Commichau
Selection-driven accumulation of suppressor mutants in bacillus subtilis: the apparent high mutation frequency of the cryptic gudB gene and the rapid clonal expansion of gudB(+) suppressors are due to growth under selection.
PLoS One: 2013, 8(6);e66120
[PubMed:23785476]
[WorldCat.org]
[DOI]
(I e)
Li-Li Chen, Jia-Le Wang, Yu Hu, Bing-Jun Qian, Xiao-Min Yao, Jing-Fang Wang, Jian-Hua Zhang
Computational design of glutamate dehydrogenase in Bacillus subtilis natto.
J Mol Model: 2013, 19(4);1919-27
[PubMed:23338837]
[WorldCat.org]
[DOI]
(I p)
Alexander K W Elsholz, Kürsad Turgay, Stephan Michalik, Bernd Hessling, Katrin Gronau, Dan Oertel, Ulrike Mäder, Jörg Bernhardt, Dörte Becher, Michael Hecker, Ulf Gerth
Global impact of protein arginine phosphorylation on the physiology of Bacillus subtilis.
Proc Natl Acad Sci U S A: 2012, 109(19);7451-6
[PubMed:22517742]
[WorldCat.org]
[DOI]
(I p)
Katrin Gunka, Stefan Tholen, Jan Gerwig, Christina Herzberg, Jörg Stülke, Fabian M Commichau
A high-frequency mutation in Bacillus subtilis: requirements for the decryptification of the gudB glutamate dehydrogenase gene.
J Bacteriol: 2012, 194(5);1036-44
[PubMed:22178973]
[WorldCat.org]
[DOI]
(I p)
Yong Heon Lee, Anthony W Kingston, John D Helmann
Glutamate dehydrogenase affects resistance to cell wall antibiotics in Bacillus subtilis.
J Bacteriol: 2012, 194(5);993-1001
[PubMed:22178969]
[WorldCat.org]
[DOI]
(I p)
Lope A Flórez, Katrin Gunka, Rafael Polanía, Stefan Tholen, Jörg Stülke
SPABBATS: A pathway-discovery method based on Boolean satisfiability that facilitates the characterization of suppressor mutants.
BMC Syst Biol: 2011, 5;5
[PubMed:21219666]
[WorldCat.org]
[DOI]
(I e)
Katrin Gunka, Joseph A Newman, Fabian M Commichau, Christina Herzberg, Cecilia Rodrigues, Lorraine Hewitt, Richard J Lewis, Jörg Stülke
Functional dissection of a trigger enzyme: mutations of the bacillus subtilis glutamate dehydrogenase RocG that affect differentially its catalytic activity and regulatory properties.
J Mol Biol: 2010, 400(4);815-27
[PubMed:20630473]
[WorldCat.org]
[DOI]
(I p)
Daniel R Zeigler, Zoltán Prágai, Sabrina Rodriguez, Bastien Chevreux, Andrea Muffler, Thomas Albert, Renyuan Bai, Markus Wyss, John B Perkins
The origins of 168, W23, and other Bacillus subtilis legacy strains.
J Bacteriol: 2008, 190(21);6983-95
[PubMed:18723616]
[WorldCat.org]
[DOI]
(I p)
Shigeki Kada, Masahiro Yabusaki, Takayuki Kaga, Hitoshi Ashida, Ken-ichi Yoshida
Identification of two major ammonia-releasing reactions involved in secondary natto fermentation.
Biosci Biotechnol Biochem: 2008, 72(7);1869-76
[PubMed:18603778]
[WorldCat.org]
[DOI]
(I p)
Fabian M Commichau, Katrin Gunka, Jens J Landmann, Jörg Stülke
Glutamate metabolism in Bacillus subtilis: gene expression and enzyme activities evolved to avoid futile cycles and to allow rapid responses to perturbations of the system.
J Bacteriol: 2008, 190(10);3557-64
[PubMed:18326565]
[WorldCat.org]
[DOI]
(I p)
Ulf Gerth, Holger Kock, Ilja Kusters, Stephan Michalik, Robert L Switzer, Michael Hecker
Clp-dependent proteolysis down-regulates central metabolic pathways in glucose-starved Bacillus subtilis.
J Bacteriol: 2008, 190(1);321-31
[PubMed:17981983]
[WorldCat.org]
[DOI]
(I p)
Fabian M Commichau, Ingrid Wacker, Jan Schleider, Hans-Matti Blencke, Irene Reif, Philipp Tripal, Jörg Stülke
Characterization of Bacillus subtilis mutants with carbon source-independent glutamate biosynthesis.
J Mol Microbiol Biotechnol: 2007, 12(1-2);106-13
[PubMed:17183217]
[WorldCat.org]
[DOI]
(P p)
B R Belitsky, A L Sonenshein
Role and regulation of Bacillus subtilis glutamate dehydrogenase genes.
J Bacteriol: 1998, 180(23);6298-305
[PubMed:9829940]
[WorldCat.org]
[DOI]
(P p)