Difference between revisions of "GerKA"
| Line 16: | Line 16: | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU03700 gerKA] | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU03700 gerKA] | ||
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http:// | + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://subtiwiki.uni-goettingen.de/interact/ ''Subt''Interact]''': [http://subtiwiki.uni-goettingen.de/interact/index.php?protein=GerKA GerKA] |
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 60 kDa, 5.206 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 60 kDa, 5.206 | ||
Revision as of 15:11, 11 November 2013
- Description: nutrient receptor
| Gene name | gerKA |
| Synonyms | |
| Essential | no |
| Product | nutrient receptor |
| Function | germination response to the combination of glucose, fructose, aspartate, and KCl |
| Gene expression levels in SubtiExpress: gerKA | |
| Interactions involving this protein in SubtInteract: GerKA | |
| MW, pI | 60 kDa, 5.206 |
| Gene length, protein length | 1632 bp, 544 aa |
| Immediate neighbours | gerKD, gerKC |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 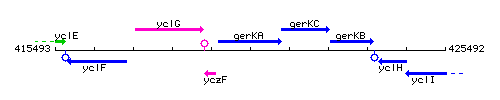
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed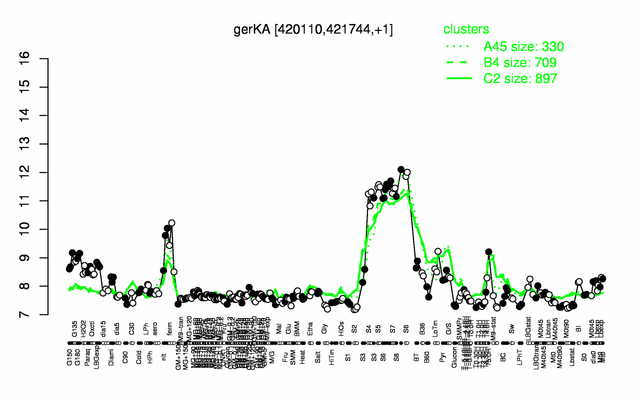
| |
Contents
Categories containing this gene/protein
germination, membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU03700
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: gerABKA family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- contains multiple membrane-spanning domains PubMed
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- outer surface of the inner spore membrane PubMed
Database entries
- Structure:
- UniProt: P49939
- KEGG entry: [3]
- E.C. number:
Additional information
- 700 molecules are present per spore PubMed
Expression and regulation
- Sigma factor: SigG PubMed
- Additional information:
- 700 molecules are present per spore PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Kerry-Ann V Stewart, Peter Setlow
Numbers of individual nutrient germinant receptors and other germination proteins in spores of Bacillus subtilis.
J Bacteriol: 2013, 195(16);3575-82
[PubMed:23749970]
[WorldCat.org]
[DOI]
(I p)
Jing-qiao Zhang, Keren K Griffiths, Ann Cowan, Peter Setlow, Ji Yu
Expression level of Bacillus subtilis germinant receptors determines the average rate but not the heterogeneity of spore germination.
J Bacteriol: 2013, 195(8);1735-40
[PubMed:23396907]
[WorldCat.org]
[DOI]
(I p)
George Korza, Peter Setlow
Topology and accessibility of germination proteins in the Bacillus subtilis spore inner membrane.
J Bacteriol: 2013, 195(7);1484-91
[PubMed:23335419]
[WorldCat.org]
[DOI]
(I p)
Sonali Ghosh, Michelle Scotland, Peter Setlow
Levels of germination proteins in dormant and superdormant spores of Bacillus subtilis.
J Bacteriol: 2012, 194(9);2221-7
[PubMed:22343299]
[WorldCat.org]
[DOI]
(I p)
Keren K Griffiths, Jingqiao Zhang, Ann E Cowan, Ji Yu, Peter Setlow
Germination proteins in the inner membrane of dormant Bacillus subtilis spores colocalize in a discrete cluster.
Mol Microbiol: 2011, 81(4);1061-77
[PubMed:21696470]
[WorldCat.org]
[DOI]
(I p)
Krzysztof Hinc, Krzysztofa Nagórska, Adam Iwanicki, Grzegorz Wegrzyn, Simone J Séror, Michal Obuchowski
Expression of genes coding for GerA and GerK spore germination receptors is dependent on the protein phosphatase PrpE.
J Bacteriol: 2006, 188(12);4373-83
[PubMed:16740944]
[WorldCat.org]
[DOI]
(P p)
Takao Igarashi, Peter Setlow
Transcription of the Bacillus subtilis gerK operon, which encodes a spore germinant receptor, and comparison with that of operons encoding other germinant receptors.
J Bacteriol: 2006, 188(11);4131-6
[PubMed:16707705]
[WorldCat.org]
[DOI]
(P p)
Swaroopa Atluri, Katerina Ragkousi, Donna E Cortezzo, Peter Setlow
Cooperativity between different nutrient receptors in germination of spores of Bacillus subtilis and reduction of this cooperativity by alterations in the GerB receptor.
J Bacteriol: 2006, 188(1);28-36
[PubMed:16352818]
[WorldCat.org]
[DOI]
(P p)