Difference between revisions of "GamA"
| Line 111: | Line 111: | ||
* '''Regulation:''' | * '''Regulation:''' | ||
** induced by N-acetylglucosamine {{PubMed|14343123}} | ** induced by N-acetylglucosamine {{PubMed|14343123}} | ||
| − | ** induced by glucosamine ([[GamR]]) {{PubMed|23667565}} | + | ** induced by glucosamine ([[GamR]]) {{PubMed|24673833,23667565}} |
* '''Regulatory mechanism:''' | * '''Regulatory mechanism:''' | ||
| − | ** [[GamR]]: transcription repression {{PubMed|23667565}} | + | ** [[GamR]]: transcription repression {{PubMed|24673833,23667565}} |
* '''Additional information:''' | * '''Additional information:''' | ||
| Line 138: | Line 138: | ||
=References= | =References= | ||
| − | <pubmed>10627040, 14343123 23667565 23876412 15755726 </pubmed> | + | <pubmed>10627040, 14343123 23667565 23876412 24673833,15755726 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 10:52, 31 March 2014
- Description: glucosamine-6-phosphate deaminase
| Gene name | gamA |
| Synonyms | ybfT |
| Essential | no |
| Product | glucosamine-6-phosphate deaminase |
| Function | glucosamine utilization |
| Gene expression levels in SubtiExpress: gamA | |
| Metabolic function and regulation of this protein in SubtiPathways: gamA | |
| MW, pI | 27 kDa, 5.793 |
| Gene length, protein length | 747 bp, 249 aa |
| Immediate neighbours | gamP, gamR |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 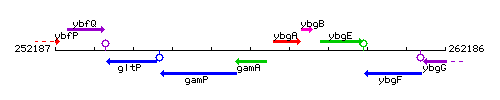
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed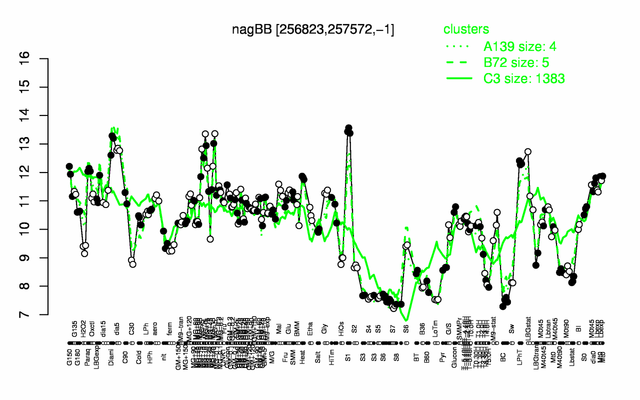
| |
Contents
Categories containing this gene/protein
cell wall degradation/ turnover, utilization of specific carbon sources
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU02360
Phenotypes of a mutant
- no growth on glucosamine PubMed
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: d-glucosamine 6-phosphate + H2O = D-fructose 6-phosphate + NH3 (according to Swiss-Prot)
- Protein family: NagB subfamily (according to Swiss-Prot)
- Paralogous protein(s): NagB
Extended information on the protein
- Kinetic information: K(M): 3.0 mM PubMed
- Modification:
- Effectors of protein activity:
Database entries
- UniProt: O31458
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Isabelle Gaugué, Jacques Oberto, Jacqueline Plumbridge
Regulation of amino sugar utilization in Bacillus subtilis by the GntR family regulators, NagR and GamR.
Mol Microbiol: 2014, 92(1);100-15
[PubMed:24673833]
[WorldCat.org]
[DOI]
(I p)
Yanfeng Liu, Long Liu, Hyun-dong Shin, Rachel R Chen, Jianghua Li, Guocheng Du, Jian Chen
Pathway engineering of Bacillus subtilis for microbial production of N-acetylglucosamine.
Metab Eng: 2013, 19;107-15
[PubMed:23876412]
[WorldCat.org]
[DOI]
(I p)
Isabelle Gaugué, Jacques Oberto, Harald Putzer, Jacqueline Plumbridge
The use of amino sugars by Bacillus subtilis: presence of a unique operon for the catabolism of glucosamine.
PLoS One: 2013, 8(5);e63025
[PubMed:23667565]
[WorldCat.org]
[DOI]
(I e)
Florence Vincent, Gideon J Davies, James A Brannigan
Structure and kinetics of a monomeric glucosamine 6-phosphate deaminase: missing link of the NagB superfamily?
J Biol Chem: 2005, 280(20);19649-55
[PubMed:15755726]
[WorldCat.org]
[DOI]
(P p)
Jonathan Reizer, Steffi Bachem, Aiala Reizer, Maryvonne Arnaud, Milton H Saier, Jörg Stülke
Novel phosphotransferase system genes revealed by genome analysis - the complete complement of PTS proteins encoded within the genome of Bacillus subtilis.
Microbiology (Reading): 1999, 145 ( Pt 12);3419-3429
[PubMed:10627040]
[WorldCat.org]
[DOI]
(P p)
C J BATES, C A PASTERNAK
FURTHER STUDIES ON THE REGULATION OF AMINO SUGAR METABOLISM IN BACILLUS SUBTILIS.
Biochem J: 1965, 96(1);147-54
[PubMed:14343123]
[WorldCat.org]
[DOI]
(P p)