FrlB
- Description: fructoselysine-6-P-glycosidase
| Gene name | frlB |
| Synonyms | yurP |
| Essential | no |
| Product | fructoselysine-6-P-glycosidase |
| Function | metabolism of aminoacylated fructose |
| Gene expression levels in SubtiExpress: frlB | |
| Metabolic function and regulation of this protein in SubtiPathways: frlB | |
| MW, pI | 36 kDa, 5.442 |
| Gene length, protein length | 984 bp, 328 aa |
| Immediate neighbours | frlO, yurQ |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 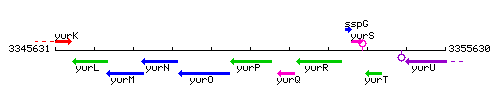
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed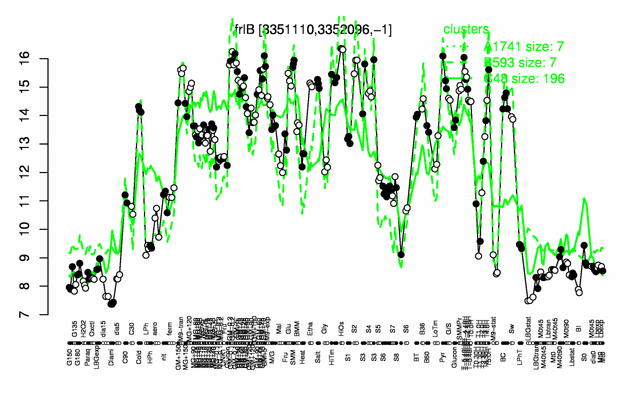
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources, utilization of nitrogen sources other than amino acids, membrane proteins, phosphoproteins, most abundant proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU32610
Phenotypes of a mutant
Database entries
- BsubCyc: BSU32610
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- phosphorylated on Arg-48 PubMed
- Effectors of protein activity:
- Localization: membrane associated PubMed
Database entries
- BsubCyc: BSU32610
- Structure: 3EUA
- UniProt: O32157
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
- the mRNA is substantially stabilized upon depletion of RNase Y PubMed
- the frlB-frlO-frlN-frlM-frlD operon is not expressed in a cshA mutant PubMed
- belongs to the 100 most abundant proteins PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium): 3695 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, exponential phase): 9758 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, early stationary phase after glucose exhaustion): 5339 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, late stationary phase after glucose exhaustion): 6442 PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Martin Lehnik-Habrink, Leonie Rempeters, Ákos T Kovács, Christoph Wrede, Claudia Baierlein, Heike Krebber, Oscar P Kuipers, Jörg Stülke
DEAD-Box RNA helicases in Bacillus subtilis have multiple functions and act independently from each other.
J Bacteriol: 2013, 195(3);534-44
[PubMed:23175651]
[WorldCat.org]
[DOI]
(I p)
Alexander K W Elsholz, Kürsad Turgay, Stephan Michalik, Bernd Hessling, Katrin Gronau, Dan Oertel, Ulrike Mäder, Jörg Bernhardt, Dörte Becher, Michael Hecker, Ulf Gerth
Global impact of protein arginine phosphorylation on the physiology of Bacillus subtilis.
Proc Natl Acad Sci U S A: 2012, 109(19);7451-6
[PubMed:22517742]
[WorldCat.org]
[DOI]
(I p)
Martin Lehnik-Habrink, Marc Schaffer, Ulrike Mäder, Christine Diethmaier, Christina Herzberg, Jörg Stülke
RNA processing in Bacillus subtilis: identification of targets of the essential RNase Y.
Mol Microbiol: 2011, 81(6);1459-73
[PubMed:21815947]
[WorldCat.org]
[DOI]
(I p)
Veronika Maria Deppe, Stephanie Klatte, Johannes Bongaerts, Karl-Heinz Maurer, Timothy O'Connell, Friedhelm Meinhardt
Genetic control of amadori product degradation in Bacillus subtilis via regulation of frlBONMD expression by FrlR.
Appl Environ Microbiol: 2011, 77(9);2839-46
[PubMed:21398478]
[WorldCat.org]
[DOI]
(I p)
Veronika Maria Deppe, Johannes Bongaerts, Timothy O'Connell, Karl-Heinz Maurer, Friedhelm Meinhardt
Enzymatic deglycation of Amadori products in bacteria: mechanisms, occurrence and physiological functions.
Appl Microbiol Biotechnol: 2011, 90(2);399-406
[PubMed:21347729]
[WorldCat.org]
[DOI]
(I p)
Hannes Hahne, Susanne Wolff, Michael Hecker, Dörte Becher
From complementarity to comprehensiveness--targeting the membrane proteome of growing Bacillus subtilis by divergent approaches.
Proteomics: 2008, 8(19);4123-36
[PubMed:18763711]
[WorldCat.org]
[DOI]
(I p)
Boris R Belitsky, Abraham L Sonenshein
Genetic and biochemical analysis of CodY-binding sites in Bacillus subtilis.
J Bacteriol: 2008, 190(4);1224-36
[PubMed:18083814]
[WorldCat.org]
[DOI]
(I p)
Elsa Wiame, Armelle Duquenne, Ghislain Delpierre, Emile Van Schaftingen
Identification of enzymes acting on alpha-glycated amino acids in Bacillus subtilis.
FEBS Lett: 2004, 577(3);469-72
[PubMed:15556630]
[WorldCat.org]
[DOI]
(P p)
Christine Eymann, Annette Dreisbach, Dirk Albrecht, Jörg Bernhardt, Dörte Becher, Sandy Gentner, Le Thi Tam, Knut Büttner, Gerrit Buurman, Christian Scharf, Simone Venz, Uwe Völker, Michael Hecker
A comprehensive proteome map of growing Bacillus subtilis cells.
Proteomics: 2004, 4(10);2849-76
[PubMed:15378759]
[WorldCat.org]
[DOI]
(P p)
Virginie Molle, Yoshiko Nakaura, Robert P Shivers, Hirotake Yamaguchi, Richard Losick, Yasutaro Fujita, Abraham L Sonenshein
Additional targets of the Bacillus subtilis global regulator CodY identified by chromatin immunoprecipitation and genome-wide transcript analysis.
J Bacteriol: 2003, 185(6);1911-22
[PubMed:12618455]
[WorldCat.org]
[DOI]
(P p)