Difference between revisions of "FadN"
(→References) |
|||
| Line 22: | Line 22: | ||
|style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || 2445 bp, 815 aa | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || 2445 bp, 815 aa | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[fadA]]'', ''[[ | + | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[fadA]]'', ''[[yuzL]]'' |
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+[EMBLCDS:CAB15273]+-newId sequences] <br/> (Barbe ''et al.'', 2009)''' | |colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+[EMBLCDS:CAB15273]+-newId sequences] <br/> (Barbe ''et al.'', 2009)''' | ||
Revision as of 10:19, 5 December 2012
- Description: 3-hydroxyacyl-CoA dehydrogenase (acetoacetyl-CoA)
| Gene name | fadN |
| Synonyms | yusL |
| Essential | no |
| Product | 3-hydroxyacyl-CoA dehydrogenase (acetoacetyl-CoA) |
| Function | fatty acid degradation |
| Gene expression levels in SubtiExpress: fadN | |
| Metabolic function and regulation of this protein in SubtiPathways: Fatty acid degradation | |
| MW, pI | 89 kDa, 6.53 |
| Gene length, protein length | 2445 bp, 815 aa |
| Immediate neighbours | fadA, yuzL |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 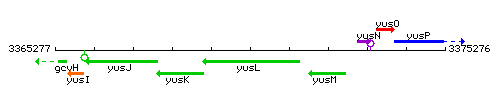
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed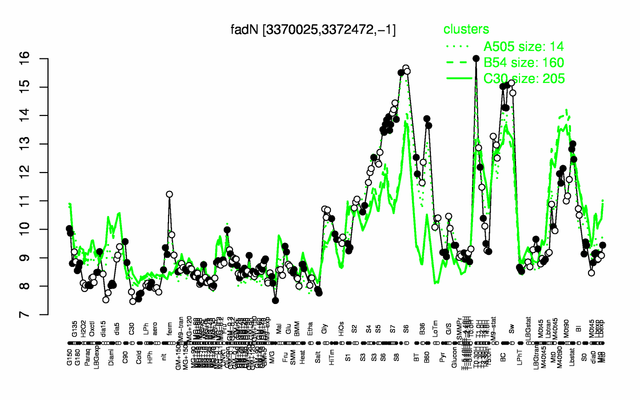
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
CcpA regulon, FadR regulon, SdpR regulon
The gene
Basic information
- Locus tag: BSU32840
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: (S)-3-hydroxyacyl-CoA + NAD+ = 3-oxoacyl-CoA + NADH (according to Swiss-Prot)
- Protein family: 3-hydroxyacyl-CoA dehydrogenase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: O32178
- KEGG entry: [3]
- E.C. number: 1.1.1.35
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Additional publications: PubMed
Yasutaro Fujita, Hiroshi Matsuoka, Kazutake Hirooka
Regulation of fatty acid metabolism in bacteria.
Mol Microbiol: 2007, 66(4);829-39
[PubMed:17919287]
[WorldCat.org]
[DOI]
(P p)
Hiroshi Matsuoka, Kazutake Hirooka, Yasutaro Fujita
Organization and function of the YsiA regulon of Bacillus subtilis involved in fatty acid degradation.
J Biol Chem: 2007, 282(8);5180-94
[PubMed:17189250]
[WorldCat.org]
[DOI]
(P p)
Hans-Matti Blencke, Georg Homuth, Holger Ludwig, Ulrike Mäder, Michael Hecker, Jörg Stülke
Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways.
Metab Eng: 2003, 5(2);133-49
[PubMed:12850135]
[WorldCat.org]
[DOI]
(P p)
José E González-Pastor, Errett C Hobbs, Richard Losick
Cannibalism by sporulating bacteria.
Science: 2003, 301(5632);510-3
[PubMed:12817086]
[WorldCat.org]
[DOI]
(I p)