CitB
- Description: trigger enzyme: aconitase and RNA binding protein
| Gene name | citB |
| Synonyms | |
| Essential | no |
| Product | trigger enzyme: aconitate hydratase (aconitase) |
| Function | TCA cycle |
| Gene expression levels in SubtiExpress: citB | |
| Interactions involving this protein in SubtInteract: CitB | |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 99 kDa, 4.903 |
| Gene length, protein length | 2727 bp, 909 aa |
| Immediate neighbours | sspO, yneN |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 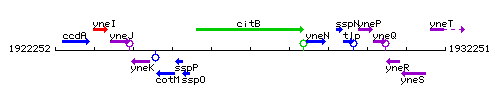
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed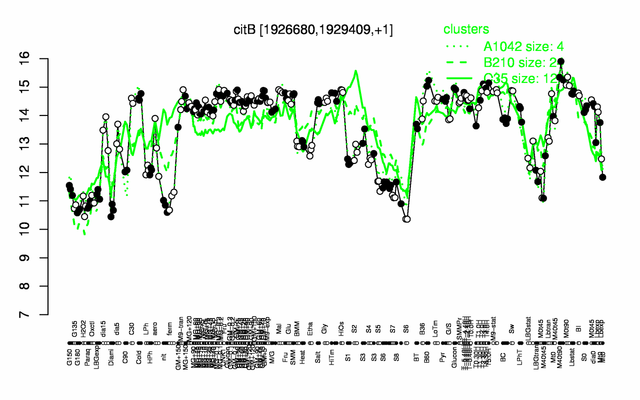
| |
Contents
Categories containing this gene/protein
carbon core metabolism, trigger enzyme, RNA binding regulators
This gene is a member of the following regulons
CcpC regulon, CodY regulon, FsrA regulon
The CitB regulon: feuA-feuB-feuC-ybbA
The gene
Basic information
- Locus tag: BSU18000
Phenotypes of a mutant
glutamate auxotrophy and a defect in sporulation PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
- A mutation was found in this gene after evolution under relaxed selection for sporulation PubMed
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Citrate = isocitrate (according to Swiss-Prot)
- Binding to iron responsive elements (IRE RNA) in the absence of the FeS cluster PubMed
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s): FeS cluster
- Effectors of protein activity:
Database entries
- Structure: 1L5J (E. coli)
- UniProt: P09339
- KEGG entry: [3]
- E.C. number: 4.2.1.3
Additional information
- B. subtilis aconitase is both an enzyme and an RNA binding protein (moonlighting protein) PubMed
- extensive information on the structure and enzymatic properties of CitB can be found at Proteopedia
Expression and regulation
- Regulation:
- repressed during growth in the presence of branched chain amino acids (CodY) PubMed
- repressed in the presence of glucose and glutamate (CcpC) PubMed
- expressed upon transition into the stationary phase (AbrB) PubMed, indirect negative regulation by AbrB PubMed
- repressed by glucose (3.7-fold) (CcpA) PubMed
- repression by glucose + arginine (CcpC) PubMed
- less expressed under conditions of extreme iron limitation (FsrA) PubMed
- part of the iron sparing response (FsrA) PubMed
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- GP683 (erm), available in Jörg Stülke's lab
- GP1441 (spc), available in Jörg Stülke's lab
- 1A999 ( citB::spec), PubMed, available at BGSC
- Expression vector:
- lacZ fusion:
- pGP700 (in pAC5), available in Jörg Stülke's lab
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Jörg Stülke's lab
- Antibody: available in Linc Sonenshein's lab
- FLAG-tag construct:
- GP1144 (spc, based on pGP1331), available in Jörg Stülke's lab
- GP1145 (kan), available in Jörg Stülke's lab
Labs working on this gene/protein
Linc Sonenshein, Tufts University, Boston, MA, USA Homepage
Jörg Stülke, University of Göttingen, Germany Homepage
Your additional remarks
References
Reviews
Karl Volz
The functional duality of iron regulatory protein 1.
Curr Opin Struct Biol: 2008, 18(1);106-11
[PubMed:18261896]
[WorldCat.org]
[DOI]
(P p)
Fabian M Commichau, Jörg Stülke
Trigger enzymes: bifunctional proteins active in metabolism and in controlling gene expression.
Mol Microbiol: 2008, 67(4);692-702
[PubMed:18086213]
[WorldCat.org]
[DOI]
(P p)
Patricia J Kiley, Helmut Beinert
The role of Fe-S proteins in sensing and regulation in bacteria.
Curr Opin Microbiol: 2003, 6(2);181-5
[PubMed:12732309]
[WorldCat.org]
[DOI]
(P p)
R L Switzer
Non-redox roles for iron-sulfur clusters in enzymes.
Biofactors: 1989, 2(2);77-86
[PubMed:2696478]
[WorldCat.org]
(P p)
Original publications
Meghna Mittal, Kieran B Pechter, Silvia Picossi, Hyun-Jin Kim, Kathryn O Kerstein, Abraham L Sonenshein
Dual role of CcpC protein in regulation of aconitase gene expression in Listeria monocytogenes and Bacillus subtilis.
Microbiology (Reading): 2013, 159(Pt 1);68-76
[PubMed:23139400]
[WorldCat.org]
[DOI]
(I p)
Gregory T Smaldone, Olga Revelles, Ahmed Gaballa, Uwe Sauer, Haike Antelmann, John D Helmann
A global investigation of the Bacillus subtilis iron-sparing response identifies major changes in metabolism.
J Bacteriol: 2012, 194(10);2594-605
[PubMed:22389480]
[WorldCat.org]
[DOI]
(I p)
Christopher T Brown, Laura K Fishwick, Binna M Chokshi, Marissa A Cuff, Jay M Jackson, Travis Oglesby, Alison T Rioux, Enrique Rodriguez, Gregory S Stupp, Austin H Trupp, James S Woollcombe-Clarke, Tracy N Wright, William J Zaragoza, Jennifer C Drew, Eric W Triplett, Wayne L Nicholson
Whole-genome sequencing and phenotypic analysis of Bacillus subtilis mutants following evolution under conditions of relaxed selection for sporulation.
Appl Environ Microbiol: 2011, 77(19);6867-77
[PubMed:21821766]
[WorldCat.org]
[DOI]
(I p)
Weihua Gao, Sen Dai, Quanli Liu, Haijin Xu, Yanlin Bai, Mingqiang Qiao
Effect of site-directed mutagenesis of citB on the expression and activity of Bacillus subtilis aconitase.
Mikrobiologiia: 2010, 79(6);774-8
[PubMed:21446632]
[WorldCat.org]
(P p)
Weihua Gao, Sen Dai, Quanli Liu, Haijin Xu, Mingqiang Qiao
CitB mutation increases the alkaline protease productivity in Bacillus subtilis.
J Gen Appl Microbiol: 2010, 56(5);403-7
[PubMed:21099137]
[WorldCat.org]
[DOI]
(P p)
Frederik M Meyer, Jan Gerwig, Elke Hammer, Christina Herzberg, Fabian M Commichau, Uwe Völker, Jörg Stülke
Physical interactions between tricarboxylic acid cycle enzymes in Bacillus subtilis: evidence for a metabolon.
Metab Eng: 2011, 13(1);18-27
[PubMed:20933603]
[WorldCat.org]
[DOI]
(I p)
Alexander G Albrecht, Daili J A Netz, Marcus Miethke, Antonio J Pierik, Olaf Burghaus, Florian Peuckert, Roland Lill, Mohamed A Marahiel
SufU is an essential iron-sulfur cluster scaffold protein in Bacillus subtilis.
J Bacteriol: 2010, 192(6);1643-51
[PubMed:20097860]
[WorldCat.org]
[DOI]
(I p)
Ahmed Gaballa, Haike Antelmann, Claudio Aguilar, Sukhjit K Khakh, Kyung-Bok Song, Gregory T Smaldone, John D Helmann
The Bacillus subtilis iron-sparing response is mediated by a Fur-regulated small RNA and three small, basic proteins.
Proc Natl Acad Sci U S A: 2008, 105(33);11927-32
[PubMed:18697947]
[WorldCat.org]
[DOI]
(I p)
Alisa W Serio, Kieran B Pechter, Abraham L Sonenshein
Bacillus subtilis aconitase is required for efficient late-sporulation gene expression.
J Bacteriol: 2006, 188(17);6396-405
[PubMed:16923907]
[WorldCat.org]
[DOI]
(P p)
Hans-Matti Blencke, Irene Reif, Fabian M Commichau, Christian Detsch, Ingrid Wacker, Holger Ludwig, Jörg Stülke
Regulation of citB expression in Bacillus subtilis: integration of multiple metabolic signals in the citrate pool and by the general nitrogen regulatory system.
Arch Microbiol: 2006, 185(2);136-46
[PubMed:16395550]
[WorldCat.org]
[DOI]
(P p)
Hans-Matti Blencke, Georg Homuth, Holger Ludwig, Ulrike Mäder, Michael Hecker, Jörg Stülke
Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways.
Metab Eng: 2003, 5(2);133-49
[PubMed:12850135]
[WorldCat.org]
[DOI]
(P p)
Hyun-Jin Kim, Sam-In Kim, Manoja Ratnayake-Lecamwasam, Kiyoshi Tachikawa, Abraham L Sonenshein, Mark Strauch
Complex regulation of the Bacillus subtilis aconitase gene.
J Bacteriol: 2003, 185(5);1672-80
[PubMed:12591885]
[WorldCat.org]
[DOI]
(P p)
C Jourlin-Castelli, N Mani, M M Nakano, A L Sonenshein
CcpC, a novel regulator of the LysR family required for glucose repression of the citB gene in Bacillus subtilis.
J Mol Biol: 2000, 295(4);865-78
[PubMed:10656796]
[WorldCat.org]
[DOI]
(P p)
C Alén, A L Sonenshein
Bacillus subtilis aconitase is an RNA-binding protein.
Proc Natl Acad Sci U S A: 1999, 96(18);10412-7
[PubMed:10468622]
[WorldCat.org]
[DOI]
(P p)
M M Nakano, P Zuber, A L Sonenshein
Anaerobic regulation of Bacillus subtilis Krebs cycle genes.
J Bacteriol: 1998, 180(13);3304-11
[PubMed:9642180]
[WorldCat.org]
[DOI]
(P p)
J E Craig, M J Ford, D C Blaydon, A L Sonenshein
A null mutation in the Bacillus subtilis aconitase gene causes a block in Spo0A-phosphate-dependent gene expression.
J Bacteriol: 1997, 179(23);7351-9
[PubMed:9393699]
[WorldCat.org]
[DOI]
(P p)
A Fouet, S F Jin, G Raffel, A L Sonenshein
Multiple regulatory sites in the Bacillus subtilis citB promoter region.
J Bacteriol: 1990, 172(9);5408-15
[PubMed:2118511]
[WorldCat.org]
[DOI]
(P p)
A Fouet, A L Sonenshein
A target for carbon source-dependent negative regulation of the citB promoter of Bacillus subtilis.
J Bacteriol: 1990, 172(2);835-44
[PubMed:2105305]
[WorldCat.org]
[DOI]
(P p)
M S Rosenkrantz, D W Dingman, A L Sonenshein
Bacillus subtilis citB gene is regulated synergistically by glucose and glutamine.
J Bacteriol: 1985, 164(1);155-64
[PubMed:2413006]
[WorldCat.org]
[DOI]
(P p)
S H Fisher, B Magasanik
2-Ketoglutarate and the regulation of aconitase and histidase formation in Bacillus subtilis.
J Bacteriol: 1984, 158(1);379-82
[PubMed:6143742]
[WorldCat.org]
[DOI]
(P p)