Difference between revisions of "CheC"
(→Expression and regulation) |
|||
| Line 112: | Line 112: | ||
=Expression and regulation= | =Expression and regulation= | ||
| − | * '''Operon:''' | + | * '''Operon:''' part of the [[fla-che operon]] |
** ''[[flgB]]-[[flgC]]-[[fliE]]-[[fliF]]-[[fliG]]-[[fliH]]-[[fliI]]-[[fliJ]]-[[ylxF]]-[[fliK]]-[[flgD]]-[[flgE]]-[[fliL]]-[[fliM]]-[[fliY]]-[[cheY]]-[[fliZ]]-[[fliP]]-[[fliQ]]-[[fliR]]-[[flhB]]-[[flhA]]-[[flhF]]-[[flhG]]-[[cheB]]-[[cheA]]-[[cheW]]-[[cheC]]-[[cheD]]-[[sigD]]-[[swrB]]'' {{PubMed|9657996,8157612,15175317}} | ** ''[[flgB]]-[[flgC]]-[[fliE]]-[[fliF]]-[[fliG]]-[[fliH]]-[[fliI]]-[[fliJ]]-[[ylxF]]-[[fliK]]-[[flgD]]-[[flgE]]-[[fliL]]-[[fliM]]-[[fliY]]-[[cheY]]-[[fliZ]]-[[fliP]]-[[fliQ]]-[[fliR]]-[[flhB]]-[[flhA]]-[[flhF]]-[[flhG]]-[[cheB]]-[[cheA]]-[[cheW]]-[[cheC]]-[[cheD]]-[[sigD]]-[[swrB]]'' {{PubMed|9657996,8157612,15175317}} | ||
** ''[[ylxF]]-[[fliK]]-[[flgD]]-[[flgE]]-[[fliL]]-[[fliM]]-[[fliY]]-[[cheY]]-[[fliZ]]-[[fliP]]-[[fliQ]]-[[fliR]]-[[flhB]]-[[flhA]]-[[flhF]]-[[flhG]]-[[cheB]]-[[cheA]]-[[cheW]]-[[cheC]]-[[cheD]]-[[sigD]]-[[swrB]]'' {{PubMed|20233303}} | ** ''[[ylxF]]-[[fliK]]-[[flgD]]-[[flgE]]-[[fliL]]-[[fliM]]-[[fliY]]-[[cheY]]-[[fliZ]]-[[fliP]]-[[fliQ]]-[[fliR]]-[[flhB]]-[[flhA]]-[[flhF]]-[[flhG]]-[[cheB]]-[[cheA]]-[[cheW]]-[[cheC]]-[[cheD]]-[[sigD]]-[[swrB]]'' {{PubMed|20233303}} | ||
| Line 122: | Line 122: | ||
** ''[[ylxF]]'': [[SigD]] {{PubMed|20233303}} | ** ''[[ylxF]]'': [[SigD]] {{PubMed|20233303}} | ||
| − | * '''Regulation:''' | + | * '''Regulation:''' see [[fla-che operon]] |
| − | |||
| − | * '''Regulatory mechanism:''' | + | * '''Regulatory mechanism:''' see [[fla-che operon]] |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
* '''Additional information:''' | * '''Additional information:''' | ||
Revision as of 14:18, 12 February 2015
- Description: control of chemotaxis by interacting with CheD, CheY-P phosphatase, inhibition of CheR-mediated methylation of MCPs
| Gene name | cheC |
| Synonyms | ylxJ |
| Essential | no |
| Product | CheY-P phosphatase |
| Function | motility and chemotaxis |
| Gene expression levels in SubtiExpress: cheC | |
| Interactions involving this protein in SubtInteract: CheC | |
| MW, pI | 22 kDa, 4.035 |
| Gene length, protein length | 627 bp, 209 aa |
| Immediate neighbours | cheW, cheD |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 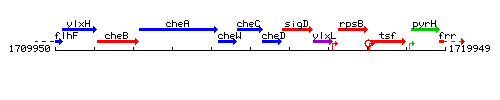
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed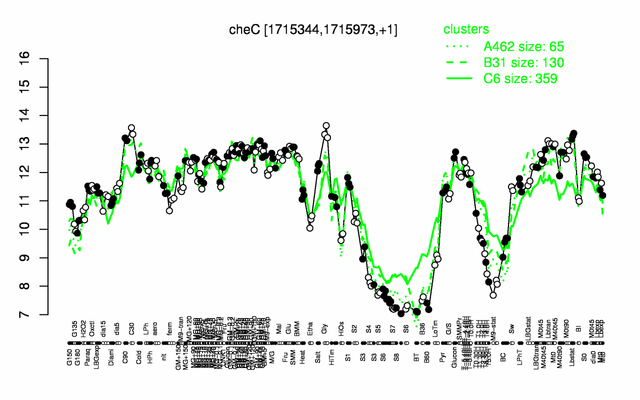
| |
Contents
Categories containing this gene/protein
protein modification, motility and chemotaxis
This gene is a member of the following regulons
CodY regulon, DegU regulon, SigD regulon, Spo0A regulon
The gene
Basic information
- Locus tag: BSU16450
Phenotypes of a mutant
Database entries
- BsubCyc: BSU16450
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: cheC family (according to Swiss-Prot)
- Paralogous protein(s):FliY
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity: interaction with CheD enhances phosphatase activity towards CheY-P PubMed
Database entries
- BsubCyc: BSU16450
- Structure:
- UniProt: P40403
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: part of the fla-che operon
- Regulation: see fla-che operon
- Regulatory mechanism: see fla-che operon
- Additional information:
Biological materials
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Serena Mordini, Cecilia Osera, Simone Marini, Francesco Scavone, Riccardo Bellazzi, Alessandro Galizzi, Cinzia Calvio
The role of SwrA, DegU and P(D3) in fla/che expression in B. subtilis.
PLoS One: 2013, 8(12);e85065
[PubMed:24386445]
[WorldCat.org]
[DOI]
(I e)
Wei Yuan, George D Glekas, George M Allen, Hanna E Walukiewicz, Christopher V Rao, George W Ordal
The importance of the interaction of CheD with CheC and the chemoreceptors compared to its enzymatic activity during chemotaxis in Bacillus subtilis.
PLoS One: 2012, 7(12);e50689
[PubMed:23226535]
[WorldCat.org]
[DOI]
(I p)
Vincent J Cannistraro, George D Glekas, Christopher V Rao, George W Ordal
Cellular stoichiometry of the chemotaxis proteins in Bacillus subtilis.
J Bacteriol: 2011, 193(13);3220-7
[PubMed:21515776]
[WorldCat.org]
[DOI]
(I p)
Ruth E Silversmith
Auxiliary phosphatases in two-component signal transduction.
Curr Opin Microbiol: 2010, 13(2);177-83
[PubMed:20133180]
[WorldCat.org]
[DOI]
(I p)
Y Pazy, M A Motaleb, M T Guarnieri, N W Charon, R Zhao, R E Silversmith
Identical phosphatase mechanisms achieved through distinct modes of binding phosphoprotein substrate.
Proc Natl Acad Sci U S A: 2010, 107(5);1924-9
[PubMed:20080618]
[WorldCat.org]
[DOI]
(I p)
Travis J Muff, George W Ordal
The diverse CheC-type phosphatases: chemotaxis and beyond.
Mol Microbiol: 2008, 70(5);1054-61
[PubMed:18990184]
[WorldCat.org]
[DOI]
(I p)
Christopher V Rao, George D Glekas, George W Ordal
The three adaptation systems of Bacillus subtilis chemotaxis.
Trends Microbiol: 2008, 16(10);480-7
[PubMed:18774298]
[WorldCat.org]
[DOI]
(P p)
Travis J Muff, George W Ordal
The CheC phosphatase regulates chemotactic adaptation through CheD.
J Biol Chem: 2007, 282(47);34120-8
[PubMed:17908686]
[WorldCat.org]
[DOI]
(P p)
Kazuo Kobayashi
Gradual activation of the response regulator DegU controls serial expression of genes for flagellum formation and biofilm formation in Bacillus subtilis.
Mol Microbiol: 2007, 66(2);395-409
[PubMed:17850253]
[WorldCat.org]
[DOI]
(P p)
Travis J Muff, Richard M Foster, Peter J Y Liu, George W Ordal
CheX in the three-phosphatase system of bacterial chemotaxis.
J Bacteriol: 2007, 189(19);7007-13
[PubMed:17675386]
[WorldCat.org]
[DOI]
(P p)
Travis J Muff, George W Ordal
Assays for CheC, FliY, and CheX as representatives of response regulator phosphatases.
Methods Enzymol: 2007, 423;336-48
[PubMed:17609139]
[WorldCat.org]
[DOI]
(P p)
Xingjuan Chao, Travis J Muff, Sang-Youn Park, Sheng Zhang, Abiola M Pollard, George W Ordal, Alexandrine M Bilwes, Brian R Crane
A receptor-modifying deamidase in complex with a signaling phosphatase reveals reciprocal regulation.
Cell: 2006, 124(3);561-71
[PubMed:16469702]
[WorldCat.org]
[DOI]
(P p)
H Werhane, P Lopez, M Mendel, M Zimmer, G W Ordal, L M Márquez-Magaña
The last gene of the fla/che operon in Bacillus subtilis, ylxL, is required for maximal sigmaD function.
J Bacteriol: 2004, 186(12);4025-9
[PubMed:15175317]
[WorldCat.org]
[DOI]
(P p)
Michael M Saulmon, Ece Karatan, George W Ordal
Effect of loss of CheC and other adaptational proteins on chemotactic behaviour in Bacillus subtilis.
Microbiology (Reading): 2004, 150(Pt 3);581-589
[PubMed:14993307]
[WorldCat.org]
[DOI]
(P p)
Hendrik Szurmant, Travis J Muff, George W Ordal
Bacillus subtilis CheC and FliY are members of a novel class of CheY-P-hydrolyzing proteins in the chemotactic signal transduction cascade.
J Biol Chem: 2004, 279(21);21787-92
[PubMed:14749334]
[WorldCat.org]
[DOI]
(P p)
Virginie Molle, Masaya Fujita, Shane T Jensen, Patrick Eichenberger, José E González-Pastor, Jun S Liu, Richard Losick
The Spo0A regulon of Bacillus subtilis.
Mol Microbiol: 2003, 50(5);1683-701
[PubMed:14651647]
[WorldCat.org]
[DOI]
(P p)
J R Kirby, C J Kristich, M M Saulmon, M A Zimmer, L F Garrity, I B Zhulin, G W Ordal
CheC is related to the family of flagellar switch proteins and acts independently from CheD to control chemotaxis in Bacillus subtilis.
Mol Microbiol: 2001, 42(3);573-85
[PubMed:11722727]
[WorldCat.org]
[DOI]
(P p)
W Estacio, S S Anna-Arriola, M Adedipe, L M Márquez-Magaña
Dual promoters are responsible for transcription initiation of the fla/che operon in Bacillus subtilis.
J Bacteriol: 1998, 180(14);3548-55
[PubMed:9657996]
[WorldCat.org]
[DOI]
(P p)
J R Kirby, C J Kristich, S L Feinberg, G W Ordal
Methanol production during chemotaxis to amino acids in Bacillus subtilis.
Mol Microbiol: 1997, 24(4);869-78
[PubMed:9194713]
[WorldCat.org]
[DOI]
(P p)
M M Rosario, G W Ordal
CheC and CheD interact to regulate methylation of Bacillus subtilis methyl-accepting chemotaxis proteins.
Mol Microbiol: 1996, 21(3);511-8
[PubMed:8866475]
[WorldCat.org]
[DOI]
(P p)
M M Rosario, J R Kirby, D A Bochar, G W Ordal
Chemotactic methylation and behavior in Bacillus subtilis: role of two unique proteins, CheC and CheD.
Biochemistry: 1995, 34(11);3823-31
[PubMed:7893679]
[WorldCat.org]
[DOI]
(P p)
L M Márquez-Magaña, M J Chamberlin
Characterization of the sigD transcription unit of Bacillus subtilis.
J Bacteriol: 1994, 176(8);2427-34
[PubMed:8157612]
[WorldCat.org]
[DOI]
(P p)