Difference between revisions of "Abn2"
| Line 20: | Line 20: | ||
|style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || 1407 bp, 469 aa | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || 1407 bp, 469 aa | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[yxiB]]'', ''[[ | + | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[yxiB]]'', ''[[yxzL]]'' |
|- | |- | ||
|style="background:#FAF8CC;" align="center"|'''[http://subtiwiki.uni-goettingen.de/yxiA_nucleotide.txt Gene sequence (+200bp) ]''' | |style="background:#FAF8CC;" align="center"|'''[http://subtiwiki.uni-goettingen.de/yxiA_nucleotide.txt Gene sequence (+200bp) ]''' | ||
Revision as of 11:55, 5 December 2012
- Description: endo-1,5-alpha-L-arabinosidase
| Gene name | abn2 |
| Synonyms | yxiA |
| Essential | no |
| Product | endo-1,5-alpha-L-arabinosidase |
| Function | arabinan degradation |
| Gene expression levels in SubtiExpress: abn2 | |
| MW, pI | 52 kDa, 7.371 |
| Gene length, protein length | 1407 bp, 469 aa |
| Immediate neighbours | yxiB, yxzL |
| Gene sequence (+200bp) | Protein sequence |
| Caution: The sequence for this gene in SubtiList contains errors | |
Genetic context 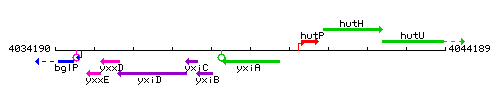
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources
This gene is a member of the following regulons
The gene
Basic information
- Locus tag:
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: glycosyl hydrolase 43 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information: With linear-alpha-1,5-l-arabinan as the preferred substrate, the enzyme exhibited an apparent K(m) of 2.0 mg ml(-1) and V(max) of 0.25 mmol min(-1) mg(-1) at pH 7.0 and 50°C. PubMed
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: extracellular (signal peptide) PubMed
Database entries
- UniProt: P42293
- KEGG entry: [3]
- E.C. number: 3.2.1.99
Additional information
Expression and regulation
- Operon: abn2 PubMed
- Regulation: repressed by glucose (4.3-fold) (CcpA) PubMed, expression is stimulated by arabinose and pectin and repressed by glucose PubMed
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Isabel de Sa-Nogueira, Lisboa, Portugal homepage
Your additional remarks
References
Birgit Voigt, Haike Antelmann, Dirk Albrecht, Armin Ehrenreich, Karl-Heinz Maurer, Stefan Evers, Gerhard Gottschalk, Jan Maarten van Dijl, Thomas Schweder, Michael Hecker
Cell physiology and protein secretion of Bacillus licheniformis compared to Bacillus subtilis.
J Mol Microbiol Biotechnol: 2009, 16(1-2);53-68
[PubMed:18957862]
[WorldCat.org]
[DOI]
(I p)
Daniele de Sanctis, Isabel Bento, José Manuel Inácio, Sónia Custódio, Isabel de Sá-Nogueira, Maria Arménia Carrondo
Overproduction, crystallization and preliminary X-ray characterization of Abn2, an endo-1,5-alpha-arabinanase from Bacillus subtilis.
Acta Crystallogr Sect F Struct Biol Cryst Commun: 2008, 64(Pt 7);636-8
[PubMed:18607095]
[WorldCat.org]
[DOI]
(I p)
José Manuel Inácio, Isabel de Sá-Nogueira
Characterization of abn2 (yxiA), encoding a Bacillus subtilis GH43 arabinanase, Abn2, and its role in arabino-polysaccharide degradation.
J Bacteriol: 2008, 190(12);4272-80
[PubMed:18408032]
[WorldCat.org]
[DOI]
(I p)
Hans-Matti Blencke, Georg Homuth, Holger Ludwig, Ulrike Mäder, Michael Hecker, Jörg Stülke
Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways.
Metab Eng: 2003, 5(2);133-49
[PubMed:12850135]
[WorldCat.org]
[DOI]
(P p)
Additional references: PubMed