XynC
- Description: endo-xylanase, preference for methylglucurono-xylan
| Gene name | xynC |
| Synonyms | ynfF |
| Essential | no |
| Product | endo-xylanase |
| Function | xylan degradation |
| Gene expression levels in SubtiExpress: xynC | |
| MW, pI | 47 kDa, 9.078 |
| Gene length, protein length | 1266 bp, 422 aa |
| Immediate neighbours | ynfE, xynD |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 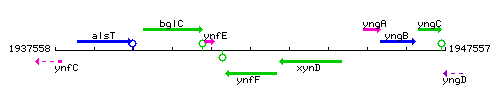
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed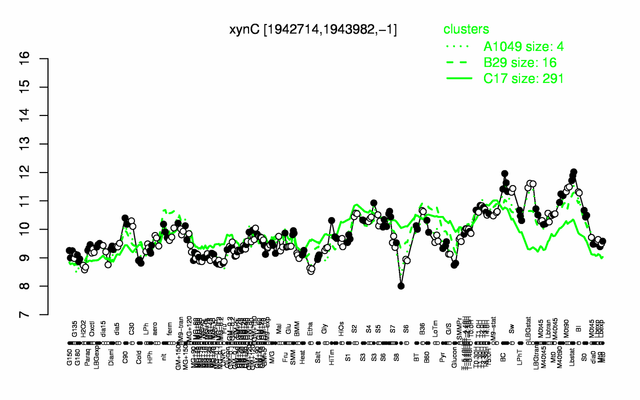
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU18150
Phenotypes of a mutant
Database entries
- BsubCyc: BSU18150
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Endohydrolysis of (1->4)-beta-D-xylosyl links in some glucuronoarabinoxylans (according to Swiss-Prot)
- Protein family: glycosyl hydrolase 30 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- extracellular (signal peptide) PubMed
Database entries
- BsubCyc: BSU18150
- UniProt: Q45070
- KEGG entry: [2]
- E.C. number: 3.2.1.136
Additional information
Expression and regulation
- Regulation:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications