Difference between revisions of "YxbA"
| Line 60: | Line 60: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU39900&redirect=T BSU39900] | ||
* '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/yxbBA-yxnB-asnH-yxaM.html] | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/yxbBA-yxnB-asnH-yxaM.html] | ||
| Line 97: | Line 98: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU39900&redirect=T BSU39900] | ||
* '''Structure:''' | * '''Structure:''' | ||
Revision as of 15:14, 2 April 2014
- Description: putative D-aspartate ligase
| Gene name | yxbA |
| Synonyms | yxaO, aslA |
| Essential | no |
| Product | putative D-aspartate ligase |
| Function | unknown |
| Gene expression levels in SubtiExpress: yxbA | |
| Metabolic function and regulation of this protein in SubtiPathways: yxbA | |
| MW, pI | 10 kDa, 4.433 |
| Gene length, protein length | 267 bp, 89 aa |
| Immediate neighbours | yxbB, yxnB |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 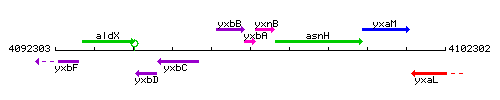
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed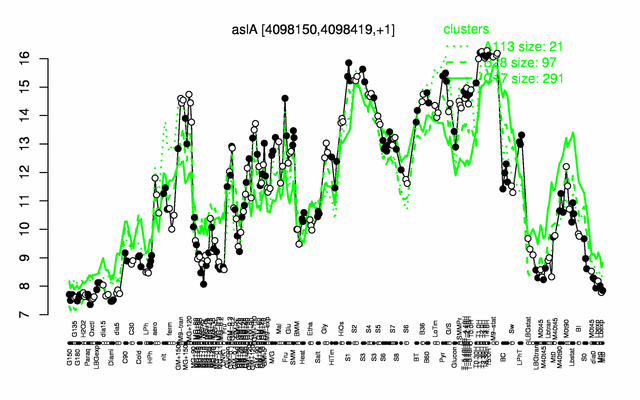
| |
Contents
Categories containing this gene/protein
poorly characterized/ putative enzymes, sporulation proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU39900
Phenotypes of a mutant
Database entries
- BsubCyc: BSU39900
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- BsubCyc: BSU39900
- Structure:
- UniProt: P46325
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Sigma factor: SigA PubMed
- Regulation:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Tetsuro Morinaga, Kazuo Kobayashi, Hitoshi Ashida, Yasutaro Fujita, Ken-Ichi Yoshida
Transcriptional regulation of the Bacillus subtilis asnH operon and role of the 5'-proximal long sequence triplication in RNA stabilization.
Microbiology (Reading): 2010, 156(Pt 6);1632-1641
[PubMed:20185509]
[WorldCat.org]
[DOI]
(I p)
Mélanie A Hamon, Nicola R Stanley, Robert A Britton, Alan D Grossman, Beth A Lazazzera
Identification of AbrB-regulated genes involved in biofilm formation by Bacillus subtilis.
Mol Microbiol: 2004, 52(3);847-60
[PubMed:15101989]
[WorldCat.org]
[DOI]
(P p)
Virginie Molle, Yoshiko Nakaura, Robert P Shivers, Hirotake Yamaguchi, Richard Losick, Yasutaro Fujita, Abraham L Sonenshein
Additional targets of the Bacillus subtilis global regulator CodY identified by chromatin immunoprecipitation and genome-wide transcript analysis.
J Bacteriol: 2003, 185(6);1911-22
[PubMed:12618455]
[WorldCat.org]
[DOI]
(P p)
Ken-Ichi Yoshida, Izumi Ishio, Eishi Nagakawa, Yoshiyuki Yamamoto, Mami Yamamoto, Yasutaro Fujita
Systematic study of gene expression and transcription organization in the gntZ-ywaA region of the Bacillus subtilis genome.
Microbiology (Reading): 2000, 146 ( Pt 3);573-579
[PubMed:10746760]
[WorldCat.org]
[DOI]
(P p)
K Yoshida, Y Fujita, S D Ehrlich
Three asparagine synthetase genes of Bacillus subtilis.
J Bacteriol: 1999, 181(19);6081-91
[PubMed:10498721]
[WorldCat.org]
[DOI]
(P p)