Difference between revisions of "YvrHb"
(→The gene) |
|||
| Line 125: | Line 125: | ||
=References= | =References= | ||
| + | <pubmed>16306698, </pubmed> | ||
# Serizawa M, Kodama K, Yamamoto H, Kobayashi K, Ogasawara N, Sekiguchi J. (2005) Functional analysis of the YvrGHb two-component system of ''Bacillus subtilis'': identification of the regulated genes by DNA microarray and northern blot analyses. ''Biosci. Biotechnol. Biochem.'' '''69:''' 2155-2169. [http://www.ncbi.nlm.nih.gov/sites/entrez/16306698 PubMed] | # Serizawa M, Kodama K, Yamamoto H, Kobayashi K, Ogasawara N, Sekiguchi J. (2005) Functional analysis of the YvrGHb two-component system of ''Bacillus subtilis'': identification of the regulated genes by DNA microarray and northern blot analyses. ''Biosci. Biotechnol. Biochem.'' '''69:''' 2155-2169. [http://www.ncbi.nlm.nih.gov/sites/entrez/16306698 PubMed] | ||
# Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | # Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | ||
Revision as of 19:47, 8 June 2009
- Description: two-component response regulator
| Gene name | yvrHb |
| Synonyms | yvrH |
| Essential | no |
| Product | two-component response regulator |
| Function | regulation cell surface maintenance |
| MW, pI | 26.6 kDa, 5.23 |
| Gene length, protein length | 714 bp, 237 amino acids |
| Immediate neighbours | yvrHa, yvrG |
| Gene sequence (+200bp) | Protein sequence |
Genetic context 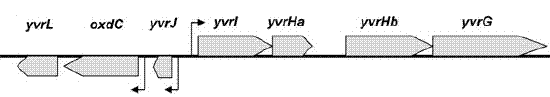
| |
Contents
The gene
Basic information
- Locus tag:
Phenotypes of a mutant
unusual autolysis and higher susceptibility to cell envelope antibiotics (bacitracin, aztreonam, cefepime, fosfomycin) PubMed
Database entries
- DBTBS entry: no entry
- SubtiList entry: no entry
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: transcription activator PubMed
- Protein family: OmpR family
- Paralogous protein(s):
The YvrHb regulon
subject to positive control: dltA-dltB-dltC-dltD-dltE, sigX-rsiX, sunA, sunT-bdbA-yolJ-bdbB, wapA-yxxG, wprA, yvrI-yvrHa PubMed
subject to negative control: lytA-lytB-lytC PubMed
Extended information on the protein
- Kinetic information:
- Domains:
- Cofactor(s):
- Effectors of protein activity:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- Structure:
- Swiss prot entry: P94504
- KEGG entry: no entry
- E.C. number:
Additional information
Expression and regulation
- Operon:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Masakuni Serizawa, Keisuke Kodama, Hiroki Yamamoto, Kazuo Kobayashi, Naotake Ogasawara, Junichi Sekiguchi
Functional analysis of the YvrGHb two-component system of Bacillus subtilis: identification of the regulated genes by DNA microarray and northern blot analyses.
Biosci Biotechnol Biochem: 2005, 69(11);2155-69
[PubMed:16306698]
[WorldCat.org]
[DOI]
(P p)
- Serizawa M, Kodama K, Yamamoto H, Kobayashi K, Ogasawara N, Sekiguchi J. (2005) Functional analysis of the YvrGHb two-component system of Bacillus subtilis: identification of the regulated genes by DNA microarray and northern blot analyses. Biosci. Biotechnol. Biochem. 69: 2155-2169. PubMed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed