Difference between revisions of "YtaG"
(→Database entries) |
|||
| Line 80: | Line 80: | ||
=== Database entries === | === Database entries === | ||
| − | * '''Structure:''' | + | * '''Structure:''' [http://www.rcsb.org/pdb/cgi/explore.cgi?pdbId=1N3B 1N3B] (from ''Escherichia coli'', 40% identity, 60% similarity) {{PubMed|12538896}} |
* '''UniProt:''' [http://www.uniprot.org/uniprot/O34932 O34932] | * '''UniProt:''' [http://www.uniprot.org/uniprot/O34932 O34932] | ||
Revision as of 13:00, 18 February 2010
- Description: dephospho-CoA kinase
| Gene name | ytaG |
| Synonyms | coaE |
| Essential | yes PubMed |
| Product | dephospho-CoA kinase |
| Function | synthesis of coenzyme A |
| Metabolic function and regulation of this protein in SubtiPathways: Coenzyme A | |
| MW, pI | 21 kDa, 4.858 |
| Gene length, protein length | 591 bp, 197 aa |
| Immediate neighbours | ytbE, ytaF |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 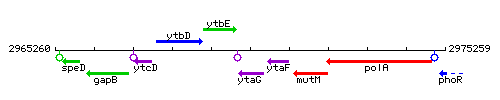
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU29060
Phenotypes of a mutant
essential PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP + 3'-dephospho-CoA = ADP + CoA (according to Swiss-Prot)
- Protein family: coaE family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification: phosphorylated on ser/ thr/ tyr PubMed
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- UniProt: O34932
- KEGG entry: [3]
- E.C. number: 2.7.1.24
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Alain Lévine, Françoise Vannier, Cédric Absalon, Lauriane Kuhn, Peter Jackson, Elaine Scrivener, Valérie Labas, Joëlle Vinh, Patrick Courtney, Jérôme Garin, Simone J Séror
Analysis of the dynamic Bacillus subtilis Ser/Thr/Tyr phosphoproteome implicated in a wide variety of cellular processes.
Proteomics: 2006, 6(7);2157-73
[PubMed:16493705]
[WorldCat.org]
[DOI]
(P p)
Alia Lapidus, Nathalie Galleron, Alexei Sorokin, S Dusko Ehrlich
Sequencing and functional annotation of the Bacillus subtilis genes in the 200 kb rrnB-dnaB region.
Microbiology (Reading): 1997, 143 ( Pt 11);3431-3441
[PubMed:9387221]
[WorldCat.org]
[DOI]
(P p)