Difference between revisions of "YrvO"
| Line 1: | Line 1: | ||
| − | * '''Description:''' | + | * '''Description:''' cysteine desulfurase<br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
| Line 10: | Line 10: | ||
|style="background:#ABCDEF;" align="center"| '''Essential''' || yes [http://www.ncbi.nlm.nih.gov/pubmed/12682299 PubMed] | |style="background:#ABCDEF;" align="center"| '''Essential''' || yes [http://www.ncbi.nlm.nih.gov/pubmed/12682299 PubMed] | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Product''' || | + | |style="background:#ABCDEF;" align="center"| '''Product''' || cysteine desulfurase |
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || unknown | |style="background:#ABCDEF;" align="center"|'''Function''' || unknown | ||
| Line 56: | Line 56: | ||
=== Additional information=== | === Additional information=== | ||
| − | |||
| − | |||
| − | |||
=The protein= | =The protein= | ||
| Line 68: | Line 65: | ||
* '''Protein family:''' | * '''Protein family:''' | ||
| − | * '''Paralogous protein(s):''' [[NifS]], [[NifZ]], [[ | + | * '''Paralogous protein(s):''' [[NifS]], [[NifZ]], [[SufS]], [[YcbU]] |
=== Extended information on the protein === | === Extended information on the protein === | ||
| Line 82: | Line 79: | ||
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| − | * '''Interactions:''' | + | * '''[[SubtInteract|Interactions]]:''' |
* '''Localization:''' | * '''Localization:''' | ||
Revision as of 09:15, 2 July 2011
- Description: cysteine desulfurase
| Gene name | yrvO |
| Synonyms | nifS |
| Essential | yes PubMed |
| Product | cysteine desulfurase |
| Function | unknown |
| MW, pI | 41 kDa, 5.463 |
| Gene length, protein length | 1137 bp, 379 aa |
| Immediate neighbours | trmU, cymR |
| Gene sequence (+200bp) | Protein sequence |
Genetic context 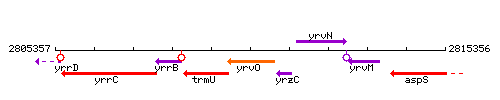
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
essential genes, poorly characterized/ putative enzymes
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU27510
Phenotypes of a mutant
essential PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
Database entries
- UniProt: O34599
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Soon-Yong Choi, Dindo Reyes, Montira Leelakriangsak, Peter Zuber
The global regulator Spx functions in the control of organosulfur metabolism in Bacillus subtilis.
J Bacteriol: 2006, 188(16);5741-51
[PubMed:16885442]
[WorldCat.org]
[DOI]
(P p)
Christine Eymann, Georg Homuth, Christian Scharf, Michael Hecker
Bacillus subtilis functional genomics: global characterization of the stringent response by proteome and transcriptome analysis.
J Bacteriol: 2002, 184(9);2500-20
[PubMed:11948165]
[WorldCat.org]
[DOI]
(P p)
J T Kaiser, T Clausen, G P Bourenkow, H D Bartunik, S Steinbacher, R Huber
Crystal structure of a NifS-like protein from Thermotoga maritima: implications for iron sulphur cluster assembly.
J Mol Biol: 2000, 297(2);451-64
[PubMed:10715213]
[WorldCat.org]
[DOI]
(P p)